Frequently Asked Questions
If you have a question that isn’t answered here, please contact us.
Are elevated levels of Protein S clinically significant?
Though not as well studied as PS deficiency, there has been shown an associated between free and total PS with cholesterol and triglycerides. Though more studies are needed, preliminary findings suggest that elevated free PS may be associated with an increased risk of ischemic heart disease.
Are the tissue factor (TF) and phospholipid (PL) compositions available?
Exact TF concentration is not available. The manufacturing process for TF/PLs is proprietary. A mixture of PL and TF is used, which form micelles of varying PL/TF concentrations and thrombin generation is then tested for release (not concentrations). Generally, TF is in the low pM range, and PLs consist of a mixture of phosphatidylcholine and phosphatidylserine.
Products: 5006010
Briefly describe the procedure for the Peviva M30 Apoptosense® assay.
Serum samples (25 µl) containing caspase-cleaved K18 (K18Asp396-NE: M30 neo-epitope) bind to an immobilized monoclonal capture antibody specific to K18. HRP-conjugated M30 antibody is added and the plates are incubated for 4 hours. Excess unbound tracer is removed by washing and TMB substrate is added. The reaction is stopped after a defined incubation period and the absorbance is measured in a microplate reader at 450 nm.
Products: P10011
Briefly describe the procedure for the Peviva M30 CytoDeath™ ELISA.
Serum samples (25 µl) containing caspase-cleaved K18 (K18Asp396-NE: M30 neo-epitope) bind to an immobilized monoclonal capture antibody specific to K18. HRP-conjugated M30 antibody is added and the plates are incubated for 4 hours. Excess unbound tracer is removed by washing and TMB substrate is added. The reaction is stopped after a defined incubation period and the absorbance is measured in a microplate reader at 450 nm.
Products: P10900
Can both serum and plasma samples be used for the M30 Apoptosense® assay?
Yes, the levels (U/L) are similar but not identical. For a particular project or series of measurements, it is recommended to use either plasma or serum for all samples investigated.
Products: P10011
Can I freeze the reagents in the Chromogenix Coatest® APC Resistance kits to prolong their stability?
Can I prepare my own controls?
Yes. However, Technoclone TGA controls are preselected for TGA use; care is taken to not induce microparticle formation. Using one’s own controls or another source might have differences in microparticles between plasmas.
Products: 5006010
Can I use Chromogenix Coatest® APC Resistance V without the predilution with FV deficient plasma? Isn't this just like using the APC R kit?
No. There is more APC in the APC V kit than in the classic kit. This makes the APC ratios twice as high among normals when not using the V def. plasma as a pre-diluent compared to the classic method. Furthermore, using the APC r-V without the pre-diluent will cause a constant change of the cut-off value due to batch-to-batch variations. This effect is not there if you use the V def. plasma, since it normalizes the plasma and ensures that the cut-off value stays the same from one batch to another.
Can I use cryoprecipitate samples with the Chromogenix Coatest® FVIII kits?
Can serum or plasma samples be used in the ELISA?
Can some components of the Peviva M30 Apoptosense® assay from different lots be mixed or combined?
No, it is strongly advised to use the components of the same lot provided with each kit only as these may affect the consistency of the results obtained.
Products: P10011
Can some components of the Peviva M30 CytoDeath™ ELISA from different lots be mixed or combined?
No, it is strongly advised to use the components of the same lot provided with each kit only as these may affect the consistency of the results obtained.
Products: P10900
Can Technothrombin® TGA be used with other species samples besides human?
Yes. However, researchers develop their own protocols and often vary the initial plasma sample dilution. For example, the dilution factor may vary depending on the species, but some researchers have had success diluting mouse plasma 1:2 in TGA buffer and dog plasma 1:3. This requires validation in the researcher’s laboratory.
Products: 5006010
Can Technothrombin® TGA measure thrombin generation of cells?
Yes. However, the experiment protocol will vary depending on the experimental goals, as thrombin generation depends on any trigger that initiates thrombin generation. In some cases, cells are used as the trigger in plasma and tested for their capacity to generate thrombin.
Products: 5006010
Can the Peviva M30 CytoDeath™ ELISA be used for all cell types?
No, the M30 CytoDeath™ ELISA can only be used for K18-positive cells (of epithelial origin) e.g. breast, lung, prostate, colon etc. Fibroblasts, lymphocytes, neuronal cells etc. can not be analyzed as these cell types do not express K18.
Products: P10900
Can the Peviva M30 CytoDeath™ ELISA be used for K18-positive cells from other species than human?
The M30 antibody recognizes caspase-cleaved keratin 18 (K18Asp396-NE M30) from human and monkey cells. K18Asp396-NE from mouse, rat, dog and xenopus cells is not detected with sufficient sensitivity. It is recommended that the presence of K18 is confirmed for the individual test cell line.
Products: P10900
Can thrombin generation of microparticles be determined?
Yes. This again will vary depending on the experimental goals as well as the source of microparticles. Trial and error may be needed in this situation. For example, cell-derived microparticles are not the same as plasma-derived microparticles. To measure thrombin generation from microparticles, isolation of the microparticles is required first, which can be completed by research protocols in the literature or using a Microparticle Filtration Unit such as the Ceveron® MFU 500.
Products: 5006010
Chromogenix Coamatic® Heparin is optimized for use with and without exogenous antithrombin. When is it recommended to add exogenous AT, and why?
It is recommended to add exogenous antithrombin when children below the age of one year are being tested. Although exogenous AT has been shown to be needless for patients with AT levels between 35-135%, pre-term newborns can have levels as low as 30%. Sufficient studies of Coamatic® Heparin have not been performed on infants, so as a precaution exogenous AT should be added. Also, for measuring heparin activities in serum, AT is needed since endogenous AT activity will be very low.
Products: K823393
Define a katal.
One katal (kat) is the amount of enzyme that converts one mole of substrate per second. Activated enzymes from Chromogenix such as FXa and thrombin are measured in nkat. 1 nkat = 1 x 10-9 mol of product released per second.
Products: S820316, S820324, S820332, S820340, S820852, S821090, S821413, S822254
Define an enzyme and a substrate.
Enzymes are proteins that catalyze most of the chemical reactions that take place in the body. The chemical compound upon which the enzyme exerts its catalytic activity is called a substrate. Proteolytic enzymes degrade their substrates, proteins and peptides, by hydrolyzing one or more peptide bond(s). For information on enzyme kinetics, see the Chromogenix Catalog or contact Diapharma Group, Inc. at info@diapharma.com.
Products: S820316, S820324, S820332, S820340, S820852, S821090, S821413, S822254
Describe chromogenic substrates in practice
Measurements made using chromogenic substrates reflect enzyme activity. Often it is more important to have knowledge about the activity of an enzyme than of the amount or mass – the quantity recorded in an immunological assay. Synthetic substrates are very sensitive, i.e. they can detect very low enzyme activities. They are in fact often more sensitive than a corresponding natural substrate.
This ability of chromogenic substrates to detect low enzyme concentrations makes them useful in, for example, the search for the presence of certain enzyme activities either in research or in quality control procedures. Sometimes there is a lack in correspondence between a natural and a chromogenic substrate in their responses to a certain enzyme preparation. For example, thrombin that has been partly degraded through autohydrolysis (ß-thrombin) reacts just as well with its chromogenic substrate as does the native form of thrombin (α-thrombin) while only native thrombin reacts with the natural substrate fibrinogen.
A chromogenic substrate is less selective, i.e. it has less discrimination in its reactivity towards related enzymes compared to the natural substrate. However, this lack of absolute selectivity can be compensated for when setting up chromogenic substrate assays. This is done by the proper selection of type of buffer, pH, relative concentrations of sample and reagents, addition of inhibitors, and/or choice of activator or incubation times. When presented with the opportunity of using one or more chromogenic substrates in a particular experimental setting for which there is no existing method, there are a few considerations that are worthwhile to make.
Substrate
If the specificity of the enzymatic activity to be measure-red is known then comprehensive overviews such as the Selectivity Tables will serve as a guide in selecting a proper substrate. The local distributor of Chromogenix products may also be contacted for advice on the choice of substrate(s). If the specificity of the enzyme is unknown, a screening procedure can be applied. When different substrates are available, such screening of the enzyme specificity can be carried out by comparing the rate of hydrolysis or pNA-generation obtained with the different substrates. Unless certain experience is available to the investigators it is usually advisable to discuss the plan and/or the result with Chromogenix. Advice on the next step can thus be given concerning either continued screening or the selection of a particular substrate that is suitable in the planned investigation.
Contaminating enzymes
If the sample to be tested with a chromogenic substrate contains more than one enzyme that may react with the same substrate, there are a number of measures that can be taken in order to eliminate the interfering/ contaminating activity. A natural or synthetic inhibitor can be introduced, the sample can be further diluted or conditions can be found (different pH and/or buffer) where the relative activities of the present enzymes are optimized. Such considerations can be based on the information below concerning temperature, pH, buffer and ionic strength.
Temperature
The rate by which the chromogenic substrate is cleaved is highly dependent on the temperature. It is therefore important to know at what temperature(s) a particular method is applicable – it may be at room (ambient) temperature, 25, 30, or 37 °C. An increase in temperature of 1 °C causes an increase in the reaction velocity of 2.5-7.5%. The temperature thus must be kept constant during the measurement and if results from different experiments are to be compared they must be performed at the same temperature. It is advisable to run the reactions in thermostated cuvettes and to use preheated stock solutions.
pH
Both Km and kcat are dependent on the pH. This means that kinetic calculations can only be made using results obtained at the same pH. Usually, the enzyme activity is measured at the pH optimum for the proteolytic activity of the enzyme. However, when several proteases are present in the same solution, as, e.g. when the sample is from plasma, it is not always advantageous to search for the pH that gives the maximum reactivity of the enzyme under investigation. Instead it is better to choose a pH where other serine proteases that may compete for the substrate have relatively lower levels of activity.
Buffers
The buffer medium and the concentration of buffer substances must be well defined. Usually Tris-HCl is used since the pKa of Tris buffer is 8.1 (25 °C), which makes it suitable for measurements at pH values between 7.3-9.3, where most of the serine proteases show maximal activities. Furthermore, this buffer is stable – it can even be autoclaved. Tris-imidazole has also been used, but is not to be recommended as imidazole is known to slightly inhibit certain proteases such as trypsin and plasmin.
Ionic strength and other additives
The appropriate ionic strength is usually obtained by adjusting the concentration of NaCl. Further substances that it may be necessary to add are CaCl2 (when Ca-dependent enzymes are studied), NaN3 (or other bactericidal agents) to prevent bacterial growth and polyethylene glycol or Tween 80 to prevent adsorption of the enzymes to the reaction vessel walls.
Substrate handling
The substrate solution is usually prepared by adding sterile water to the dry powder. Chromogenic substrates with low solubility in water can be dissolved in DMSO (dimethyl sulfoxide) and then diluted in water. The final DMSO concentration should preferably not exceed 10% in the reaction mixture. Chromogenic substrates dissolved in sterile water are stable for more than 6 months in the refrigerator (2 – 8 °C) and for several weeks at room temperature (25 °C). The stability is considerably reduced in alkaline buffers. Furthermore, contamination by microorganisms and exposure to light for longer periods should be avoided. The substrate concentration should be chosen so that linear kinetics is obtained. A substrate concentration of twice the Km (2 x Km ) is usually appropriate.
Describe heparin's interaction with antithrombin. Why is the anti-FXa method a better way to measure heparin activity?
Slow protease-antithrombin interactions are enhanced dramatically in the presence of certain sulfated polysaccharides like heparan sulfate. Heparin is a commercial preparation of heparan sulfate, and binds antithrombin, the major inhibitor of coagulation in plasma and thrombin, thereby catalyzing the thrombin-AT reaction. Binding to antithrombin induces a conformational change in AT that facilitates its reaction with thrombin. Thrombin binds to heparin in a non-specific manner and slides along the chain until it encounters the bound AT. The affinity of heparin to the thrombin-AT (TAT) complex is much lower than to free AT. Heparin will therefore dissociate from the TAT complex, which is rapidly removed from the blood circulation by the liver and the result is a stable protease inhibitor complex, which is rapidly removed and catabolized. The anti-FXa assays are more specific since they measure the ability of heparin-accelerated antithrombin to inhibit a single enzyme. Either plasma or purified AT can be used. More precise determination of unfractionated heparin and low molecular weight heparin are possible.
Describe the location and function of the various phospholipids.
Cardiolipin (CL), phosphatidylserine (PS), phosphatidylethanolamine (PE), phosphatidylcholine (PC), and phosphatidylinsitol (PI), Cardiolipin (diphosphatidylglycerol) (CL) – located in the inner mitochondrial membrane. It has an anionic charge and the function is unknown, as cardiolipin is not involved in the immune or coagulation systems.
Phosphatidylserine (PS) – located in the inner platelet membrane and, during activation, the exterior platelet membrane. It carries an anionic charge, and is the primary phospholipid which promotes the anticoagulant protein C pathway, providing feedback inhibition of thrombin formation.
Phosphatidylethanolamine (PE) – located in both the interior and exterior of cell membranes. It carries a neutral (zwitterionic) charge, and promotes the anticoagulant protein C pathway, but to a lesser degree than PS.
Phosphatidylcholine (PC) – located in the interior and exterior of cell membranes. It carries a neutral charge, and promotes the anticoagulant protein C pathway, but to a lesser degree than PS.
Phosphatidylinsitol (PI) – located in the interior and exterior of cell membranes. It carries a cationic charge, and promotes the anticoagulant protein C pathway, but to a lesser degree than PS.
Describe the protein C pathway and APC resistance. What is the relative risk of venous thrombosis for those who are APC resistant?
Protein C is a vitamin-K-dependent glycoprotein and plasma proenzyme of a serine protease that plays a key role in the down-regulation of blood coagulation. It is activated in vivo by the thrombin-thrombomodulin complex on the surface of intact endothelial cells. Activated protein C (APC) functions as a circulating anticoagulant through proteolytic cleavage and inactivation of the coagulation factors Va and VIIIa. The cleavage occurs at three sites in the heavy chain of each protein. The anticoagulant activity of APC is potentiated by the free form of Protein S (about 60% of PS in plasma is bound to C4bBP, and 40% is in free form) and FV. APC Resistance is actually due to a defect in the protein C pathway, in the factor V molecule as opposed to the activated protein C molecule. APC Resistance is an autosomal dominant hereditary defect mainly due to a point mutation resulting in an amino acid change in the FV gene (Ag506 to Gln mutation, or Factor V Leiden mutation). The mutation destroys one of the three cleavage sites, rendering FVa partially resistant to APC-mediated degradation. APC resistance occurs in 3-5% of the general population, but varies largely in different parts of the world. Up to 90% of APC resistance cases are due to the Factor V:Q506 gene mutation. The relative risk of DVT for carriers of the FV:Q506 mutation is estimated to be 8-fold for heterozygotes and 80-fold for homozygotes.
Do you have applications for the DiaPharma Factor X kit on automated coagulation analyzers?
Yes, we have applications for the ACL Advance, AMAX, BCS, STA, STA Compact, STA-R, and SysmexCA 1500, as well as for microplates. Our applications library is growing, so please check with DiaPharma for the latest updates.
Products: KDPGFX
Does hemolysis in a sample interfere with Peviva M30 Apoptosense® measurements?
The M30 Apoptosense® assay is NOT sensitive to highly elevated hemoglobin levels, which means that haemolized samples can be assayed although this is not recommended. The results obtained with the M30 Apoptosense® assay are also NOT affected by Heparin < 10 U/ml.
Products: P10011
Does the Chromogenix Coatest® SP FVIII detect animal plasma?
Does the M30 CytoDeath™ ELISA also detect necrosis?
No. Due to the specificity of the M30 antibody to a caspase cleavage product, the M30 CytoDeath™ ELISA recognizes apoptotic cells only, not necrotic or viable cells.
Products: P10900
Does the Peviva M30 Apoptosense® assay measure caspase activity?
No, the M30 Apoptosense® assay does not measure caspase activity. The assay measures the amount of accumulated caspase cleavage product, which provides superior sensitivity and a wider analytical window.
Products: P10011
Explain the function of t-PA.
tPA activates plasminogen to plasmin, and also binds to fibrin. It is synthesized mainly in vascular endothelial cell and is secreted into the plasma continuously, and also through the acute release of tPA. tPA is a relatively poor activator of plasminogen in the absence of fibrin due to the low affinity of tPA for its substrate. However, tPA has a high affinity for fibrin, and binding increases its activating capacity up to 1000 fold. This dramatic increase is attributed to specific binding sites on the fibrin surface that concentrate and correctly orientate tPA with its substrate, as well as promote efficient clot lysis.
Explain the types of protein C deficiency, and the clinical manifestations.
Hereditary protein C deficiency is inherited as an autosomal dominant trait. Heterozygotes for protein C deficiency have protein C activity or antigen levels of 30-70% normal, whereas homozygotes with a severe defect have levels below 1%. The prevalence of protein C deficiency is 2-5% in patients with thromboembolic disease. Two types of protein C deficiency states are recognized. In type I deficiency, which is the most common type of disorder, the plasma concentration of protein C is reduced both in functional and immunological assays. This reflects a genetic defect causing a reduced biosynthesis of protein C. Type II deficiency is characterized by normal protein C antigen levels, but with decreased functional activity. This type of defect reflects synthesis of abnormal molecules with reduced function. The most common clinical manifestation of symptomatic heterozygous protein C deficiency is deep vein thrombosis (DVT) of the lower extremities. Patients with homozygous protein C deficiency usually suffer from severe and fatal thrombosis in the early stage of life.
Protein C deficiency can also be acquired. Protein C level is influenced by various diseases and drugs such as DIC, DVT, liver disease, sepsis, oral anticoagulant therapy, and surgery.
In contrast, elevated Protein C levels have been reported in such cases as diabetic patients and with the use of anabolic steroids and oral contraceptives. Elevated levels of protein C have no known clinical significance.
Factor X or anti-Xa Assay. Which Do I Use?
Factor X or anti-Xa Assay. Which Do I Use?
David L. McGlasson, MS, MLS(ASCP)cm, GA Fritsma, MS, MLS(ASCP)cm
59th Clinical Research Division
JBSA Lackland, TX
How do factor V and VIII levels affect the Chromogenix Coatest® APC Resistance tests?
A study by Colucci et al. (Thromb. Haemost. 1994, 72:987 – 988) using the classical APTT method (no FV-deficient plasma pre-dilution) showed that changes in FV levels between 12.5-100% did not modify the response to APC. Other experience has shown that the Coatest® APC Resistance V test may provide ratio values approximately 0.3 units below the median level, which can be fairly close to the cut-off value, when FV levels are 0-40%. This may be explained by the fact that although there is often no abnormal bleeding tendency in heterzygotes with factor V deficiency, prolonged PT and APTT times are observed (ref: Sartori et al. Familial association of hypoplasminogenemia and heterozygous factor V deficiency. Clin Appl Thromb Hemost. 1999; 5(4): 277-281; Salooja N et al. Severe factor V deficiency and neonatal intracranial haemorrhage: a case report. Haemophilia. 2000; 6(1): 44-46. One might also see a slightly decreased ratio in patients with severe liver disease.
According to Chromogenix, FVIII samples above 1.8 IU/ml may result in a reduction of the APC ratio of approximately 0.2 units, although the actual correlation between FVIII activity and APC ratio appears to be weak. When sampling, therefore, the patient should be at rest in order to decrease the FVIII level due to stress.
How do I know mucus samples are not cross-contaminated by a nearby fish when I am collecting my samples?
It is recommended that mucus under the gill be swabbed, if cross contamination from nearby fish is a concern. This can be a concern with net fishing, where high- and low- level vitellogenin fish are in close proximity to each other.
Products: TE1037, TE1035, TE1034, TE1042, TE1040, TE1043, TE1046, TE1049, TE1039
How do I know mucus samples are not cross-contaminated with other environmental substances that may interfere with the assay and affect my results?
The antibodies used in the assays are very specific and do not recognize other components. Furthermore, the extraction buffer (500μL) used to dilute the mucus collected with the swab has a dilution effect that will dilute out a rare occurrence of an environmental contaminant. In development of the assay, aquarium water was used as a negative control and no contamination from the mucus samples was detected in comparison to the aquarium water.
Products: TE1037, TE1035, TE1034, TE1042, TE1040, TE1043, TE1046, TE1049, TE1039
How do Protein S and APC interact?
Only the free, native form of PS binds to APC and functions as a cofactor. PS has the highest affinity for negatively charged phospholipids of all the vitamin K-dependent proteins and has been shown to increase by approximately 10-fold, the affinity of APC for membranes or vesicles containing such phospholipids. This may be of physiological importance since APC degrades preferentially membrane-bound FVa and FVIIIa, but not the circulating, inactivated co-factors. In addition to increasing the affinity of APC to membranes, PS also enhances the cleavage of FVa by APC and works in concert with FV to increase the ability of APC to inactivate FVIIIa.
How do the pharmacokinetics of LMW heparins differ from UF heparin, and what are the therapeutic ranges for each?
When injected subcutaneously, the bioavailability of UF ranges from 10-90%, whereas the bioavailability of LMW heparin is greater than 90% and is independent of dose. LMW heparins exhibit much less binding to plasma proteins than UF heparin, and do not accumulate in the liver or spleen, giving them a longer plasma half-life. The dose-response curve of LMW heparin also tends to be linear.
The therapeutic range for UF heparin is 0.3-0.7 IU/ml, while the range for LMW heparin is less clearly defined. Some clinicians maintain that is 0.4-1.1 IU/ml, or more conservatively, 0.5-1.0 IU/ml (anti-FXa method).
How does a specific gene polymorphism, PAI-1 4G genotype, relate to CVD risk?
For a detailed description of the 4G polymorphism, read Kohler et al. PAI-1 and Coronary Artery Disease. NEJM 2000; 342 (24): 1792-1801. In some studies, the 4G allele (four guanine bases) was significantly associated with high plasma PAI-1 concentrations, and was most strongly associated with previous MI, as well as risk of future MI. Studies have shown that subjects who are homozygous for the 4G allele have plasma PAI-1 concentrations approximately 25% higher than those with the 5G allele (5 guanine bases). Similarly, among patients with hypertriglyceridemia, those with the 4G allele also have higher plasma PAI-1 concentrations than those with the 5G allele. There are still conflicting data on the strength of the relation between PAI-1 gene polymorphism and MI, but it is suggested that the 4G allele is more likely to contribute to MI, particularly in the presence of hypertriglyceridemia.
How does a syphilis infection affect the REAADS anti-cardiolipin test?
Patients with current or prior syphilis infections may have a positive result without increased risk of thrombosis. Anti-cardiolipin antibodies can appear transiently at low levels during many infections. If a patient first tests positive while there are clinical signs of infection, the test should be repeated after an interval of six months.
How does heparin interact with antithrombin? Why is the anti-FXa method a better way to measure heparin activity?
Slow protease-antithrombin interactions are enhanced dramatically in the presence of certain sulfated polysaccharides like heparan sulfate. Heparin is a commercial preparation of heparan sulfate, and binds antithrombin, the major inhibitor of coagulation in plasma and thrombin, thereby catalyzing the thrombin-AT reaction. Binding to antithrombin induces a conformational change in AT that facilitates its reaction with thrombin. Thrombin binds to heparin in a non-specific manner and slides along the chain until it encounters the bound AT. The affinity of heparin to the thrombin-AT (TAT) complex is much lower than to free AT. Heparin will therefore dissociate from the TAT complex, which is rapidly removed from the blood circulation by the liver and the result is a stable protease inhibitor complex, which is rapidly removed and catabolized. The anti-FXa assays are more specific since they measure the ability of heparin-accelerated antithrombin to inhibit a single enzyme. Either plasma or purified AT can be used. More precise determination of unfractionated heparin and low molecular weight heparin are possible.
How is functional activity of protein S tested?
Many labs perform functional clotting assays to test free PS activity. These will detect PS deficiency, but will not discriminate between Types I, II, and III. Clotting tests utilize an excess of protein S deficient plasma, thereby keeping the amount of prothrombin essentially constant. APTT and PT methods commonly used. In a FXa based method, coagulation is triggered by FXa in the presence of calcium ions and phospholipids. This method is not currently used in routine settings, however.
How is plasminogen converted to plasmin?
Plasminogen is activated by endogenous, natural activators, and exogenous activators. Activation by its natural activators, tPA and uPA, involves a bond cleavage at a specific site in the plg molecule, which gives rise to a two-chain molecule linked by two disulfide bonds. tPA binding to fibrin concentrates and correctly orientates the tPA and plasminogen, as well as inducing a conformational changes in the molecules that promote efficient clot lysis. uPA can only activate plasminogen in the presence of fibrin, but it does not bind to, nor is it activated by fibrin. Streptokinase is an exogenous plasminogen activator and functions by forming a 1:1 complex with human plg. This complex can function as an activator of other plg molecules. Complex formation is accompanied by a conformational change in the plg molecule, exposing the active site.
How long is a sample viable for testing?
A sample is tested 1 hour after draw to 3 hours after draw. The manufacturer is currently evaluating sample testing times less than 1 hour after draw.
How should samples for the Peviva M30 CytoDeath™ ELISA be stored?
Cell extracts should be frozen at -20°C immediately, if not used within 4 hours.
Products: P10900
How should serum or plasma samples for the Peviva M30 Apoptosense® assay be stored?
Serum and plasma samples should be stored on ice or at +4 before freezing at -80 °C.
Products: P10011
How soon after exposure will the VTG protein be detected in mucus compared to blood/homogenate and compared to the detection of vtg gene expression?
How specific is the Peviva M30 CytoDeath™ ELISA for apoptosis?
Some methods, such as TUNEL (labeling of double-stranded DNA breaks) do not distinguish between necrotic and apoptotic cells. The M30 neo-epitope on keratin 18 is only exposed after caspase cleavage occurring during apoptosis.
Products: P10900
I need an assay that complies with the USP monograph for the determination of heparin activity. What test kits are suitable?
The USP states that the activity of heparin sodium and heparin calcium should be determined by both a clotting assay and a chromogenic assay. The chromogenic assay consists essentially in the measurement of the anti-FXa activity of the test preparation against the USP Heparin Sodium Reference Standard. All of the chromogenix heparin kits meet this specification. Antithrombin, FXa, and the chromogenic substrates from Chromogenix are suitable for the USP guidelines. The anti-FXa assays are more specific since they measure the ability of heparin-accelerated antithrombin to inhibit a single enzyme. Either plasma or purified AT can be used. More precise determination of unfractionated heparin and low molecular weight heparin are possible.
If a patient plasma is hemolytic or lipemic, how should I analyze it?
Run a blank and subtract it from your sample values. The sample blank activity is determined following the method procedure by substituting the plasmin working solution with a mixture containing 1 volume of plasmin solvent and 1.5 volumes of working buffer.
If I want to determine FX in plasma, but do not want to buy a kit, what other options are there?
S-2337™ was previously included in the now discontinued Chromogenix Coatest® FX kit, so another substrate must be employed. S-2765™, which is also a substrate for FXa, can therefore be used, and a method based on the activation of FX in the presence of calcium using Russell’s Viper Venom (RVV) as the activator is described in the Chromogenix catalog. DiaPharma has composed a new DiaPharma Factor X kit based on S-2765™.
If thrombin is formed, will it hydrolyze the substrate S-2765™ when using Chromogenix Coamatic®, Coatest® SP, or Coatest® SP4 FVIII kits?
In the Chromogenix Coatest® Heparin package insert, the section Limitations of the procedure states that in some pathological states, plasma alone my hydrolyze S-2222™, and that to determine the interference one should substitute FXa with an equal volume of buffer...Why is this necessary?
In some clinical situations such as sepsis, DIC, and cancer, the plasma itself could contain enzymes that might be able to hydrolyze S-2222™. If that happened, background activity could be seen and would therefore lead to an underestimation of the heparin level in the sample. To rule out background activity in the sample, the assay can be run without FXa and instead, and equal volume of buffer is substituted. The absorbance obtained is then subtracted from the absorbance from the normal run.
Is administering one Anti-Phospholipid test sufficient for diagnosis?
No, it has been decided that more than one test is should be performed. For example, a coagulation assay for the detection of LA and an ELISA for the detection of anti-phospholipid antibodies or cofactors like anti-CL or anti-b2GPI are recommended. Future testing may require a panel of tests. Physicians should interpret the results of the APS tests in light of the patient’s history, physical findings, and other diagnostic procedures.
Is it necessary to correct mucus (or blood, WBH) VTG against the protein alone?
Is it OK to use glass tubes for the Chromogenix Factor VIII assay?
Is the Peviva M30 Apoptosis® assay sensitive and robust?
The M30 Apoptosense® assay is very sensitive. The stability of the detected K18Asp396-NE M30 neo-epitope makes the assay very robust and reproducible.
Products: P10011
Is the Peviva M30 CytoDeath™ ELISA sensitive and robust?
The assay is very sensitive, as it easily detects apoptosis in less than 5 000 carcinoma cells. The stability of the detected K18Asp396-NE M30 neo-epitope makes the assay very robust and reproducible. The established HRP-TMB ELISA method is very reproducible compared to e.g. caspase activity assays that require an optimal reaction buffer, and are sensitive to changes in temperature and substrate concentration.
Products: P10900
Is there an ultrasensitive Perch ELISA available? Are there any suggestions for modifying the Perch ELISA if lower VTG values need to be detected in mucus?
The TECO® Perch Vitellogenin ELISA is the only commercially available ELISA for Perch/Perciformes. At the moment, there is not enough demand for an ultra-sensitive protocol. One option using the current protocol is to lower the dilution factor from 1:10 (50μL sample + 450μL extraction buffer) to 1:6 (50μL sample + 250μL extraction buffer). In development, this has been shown to increase the sensitivity of the assay appropriately.
Products: TE1037, TE1035, TE1034, TE1042, TE1040, TE1043, TE1046, TE1049, TE1039
Is there any influence from alpha2-macroglobulin? If so, how can it be overcome?
Coamatic® Plasmin Inhibitor (discontinued) is not influenced by alpha2-macroglobulin up to 9 mmol/l, which is approximately three times higher than the normal level in human plasma.
Kit Symbols
 |
 |
 |
 |
| Reference Number | Lot Number | In Vitro Diagnostic | Intended Use |
 |
 |
 |
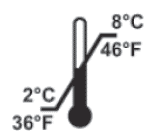 |
| Sigma (“N” tests) | Instructions | Expiration Date | Storage Temperature |
 |
 |
 |
 |
| CE Mark | Caution | Caution | Manufacturer |
MCF-7 cells are known to be defective in caspase-3 activity. Can I still use the Peviva M30 CytoDeath™ ELISA?
Yes, M30 CytoDeath™ ELISA can be used in cells lacking active caspase-3 as also other caspases, but no other proteases can generate the M30 neo-epitope. Apoptotic MCF-7 cells generate M30-activity due to activity of those other caspases.
Products: P10900
Protein concentrations in plasma
| Component | Molecular Weight kDa | Plasma Concentration mg/l | Plasma Concentration μmol/l |
|---|---|---|---|
| Fibrinogen | 330 | 3000 | 9 |
| Prothrombin | 72 | 150 | 2 |
| Factor V | 330 | 20 | 0.05 |
| Factor VII | 50 | 0.5 | 0.01 |
| Factor VIII | 330 | 0.1 | 0.0003 |
| Factor IX | 56 | 5 | 0.09 |
| Factor X | 59 | 8 | 0.13 |
| Factor XI | 160 | 5 | 0.03 |
| Factor XII | 80 | 30 | 0.4 |
| Factor XIII | 320 | 10 | 0.03 |
| Protein C | 62 | 4 | 0.06 |
| Protein S | 70 | 10 (free) | 0.14 |
| Protein Z | 62 | 2 | 0.03 |
| Prekallikrein | 86 | 50 | 0.6 |
| HMW kininogen | 120 | 70 | 0.6 |
| Fibronectin | 450 | 300 | 0.7 |
| Plasminogen | 92 | 200 | 2 |
| t-PA | 60 | 0.005 | 0.0001 |
| Urokinase | 53 | 0.004 | 0.0001 |
| Antithrombin | 58 | 145 | 2.5 |
| Heparin Cofactor II | 66 | 80 | 1.2 |
| Plasmin Inhibitor | 63 | 60 | 1 |
| Protein C Inhibitor | 57 | 4 | 0.07 |
| α2-Macroglobulin | 725 | 2000 | 3 |
Products: 5006010, 5344101, K036-001, K051-001, K255539, K822585, K823393, K824086, K824094, KDPGFX, S820316, S820324, S820332, S820340, S820852, S821090, S821413, S822254, TC12007, TC12080, TC16000, 800070
Some methods measure the late stages of apoptosis. What is known for the Peviva M30 CytoDeath™ ELISA?
The M30 CytoDeath™ ELISA detects apoptosis early on, as opposed to methods that measure DNA fragmentation.
Products: P10900
Sometimes when I run the Chromogenix Coamatic® FVIII assay, I see an upward drift in my activity from the first to the last samples. Why is this?
Thrombin, 1 NIH unit, is included in the factor reagent. It has been shown that thrombin can activate FX, and over time you will see an increase of up to 5% activity. If you are doing numerous samples at once on an automated instrument, you might want to consider Coatest® FVIII.
Substrate kinetic and selectivity tables
Kinetic data for the chromogenic substrates available from Chromogenix. Suitable chromogenic substrates are listed for a number of serine proteases, most of them part of the cascade systems in blood. Some of the substrates are cleaved by more than one enzyme although at different rates. The kinetic analyses of the enzymatic cleavage of pNA from the substrates were performed under strictly standardized conditions using the clinical chemistry analyzer Cobas Mira S.
A stable, well-defined temperature is vital for all enzyme kinetic studies and in this study all reactions were performed at 37°C. A suitable buffer was chosen for each enzyme and the pH value given in the compilation is the value to which it was adjusted at 25°C. Note that the pH value of Tris buffers decreases as the temperature increases, at the rate of approximately 0.1 unit per °C (50 mM Tris-HCl). The kinetics of the reaction was followed spectrophotometrically by measuring the change in absorbance over time, ΔA/min. To ensure the highest precision, ΔA/min was measured at four different substrate concentrations. Insertion of the ΔA/min values into Eq. 18, followed by linear regression gave Km, kcat and Vmax for the reaction.
Summarize the b2-Glycoprotein 1 test kit. What is the normal range?
Patient serum is diluted with sample diluent and incubated in microwells coated with human b2GPI. Antibodies to b2GPI present in the sample will bind to the coated wells. After washing, enzyme conjugated anti-human IgG, IgM, or IgA immunoglobulin is added, the wells are washed again, substrate added, and color development measured in a spectrophotometer at 450 nm. With a simple calculation, semi-quantitative results in G, M, or A units are available in less than 1 hour. The normal range is less than 20 units for each isotype (IgG, IgM, or IgA).
Summarize the IgG/IgM and IgA REAADS anti-cardiolipin test. What is the normal range?
The test is an indirect ELISA. Diluted serum samples, calibrator sera, and controls are incubated in cardiolipin coated microwells, allowing aCL antibodies present in the samples to react with the immobilized antigen. After their removal of unbound serum proteins by washing, antibodies specific for human IgG, IgM, or IgA labeled with HRP are added forming complexes with the CL bound antibodies. Following another washing step, the bound enzyme-antibody conjugate is assayed by the addition of TMB and H2O2 as the chromogenic substrate. Color develops in the wells at an intensity proportional to the serum concentration of aCL antibodies. The OD is read at 450 nm.
Normal ranges are less than 23 GPL (IgG per liter), less than 11 MPL (IgM per liter), and 22 APL (IgA per liter).
The Chromogenix Factor Xa reagent says it is 71 nkat, but I need to know what that is in g/ml and mol/ml.
One katal (kat) is the amount of enzyme that converts one mole of substrate per second. Activated enzymes from Chromogenix such as FXa and thrombin are measured in nkat. 1 nkat = 1 x 10-9 mol of product released per second. The conversion is as follows:
FXa has a MW approximately 44,000.
The specific activity for FXa is 1.9 nkat/mg, as determined with chromogenic substrate S-2222™.
This gives that 71 nkat corresponds to 37.4 mg FXa (=37.4 x10-6 g FXa)
This corresponds to 8.5 x 10-10 mol = 0.85nmol.
The concentration in mol/l and g/l will depend on the dilution volume you choose.
Note: 1 IU corresponds to 20 nkats, which means 71 nkatS-2222 = 3.55 IU.
The dilution of the 0.1 IU/ml heparin solution in Chromogenix Coatest® Heparin can cause some confusion. Please explain the Coatest® Heparin standardization.
Keep in mind the final concentration of heparin is 0.1 IU/ml plasma. Following the package insert, there is a known concentration of 0.1 IU/ml heparin in the standard, regardless of the dilution, provided the samples are all treated the same way. 100 ml of the 0.1 IU/ml heparin dilution is equivalent to a heparin concentration of 0.1 IU/ml plasma in the assay. This concentration should be compared to the patient-sample. If the analyst takes 100 ml of the heparin dilution (0.1 IU/ml) and dilutes it 1:10, it is comparable to 100 ml of the patient plasma also diluted in the same way. If the analyst get the same absorbance for the sample as for the standard, it contains the same amount of heparin. The important thing is the analyst must always treat patient samples in the same manner as the standards.
Products: K255539
The majority of the Chromogenix substrate library has an Arginine (Arg or R) group at the P1 position (the amino acid position that occurs at the preferred cleavage site). Why is this?
The Chromogenix line is geared toward the proteins involved in hemostasis. These are a group of proteolytic enzymes that comprise the serine proteases, which cleave mainly at the C-terminal side of the basic amino acids arginine or lysine. The peptides at the P2, P3, and P4 positions contribute to the substrate’s specificity. Note that the substrates for plasmin cleave at a lysine group. Other protease groups are aspartic proteases (like pepsin), metallo proteases, and cysteine proteases (which include caspases, with an asp cleavage site).
Products: S820316, S820324, S820332, S820340, S820852, S821090, S821413, S822254
The standard pre-dilutions for the Coatest® SP FVIII assay dont make sense to me. For example, to get 100%, I dilute 100 ml plasma with 50 ml buffer, but to get my 50% standard, I dilute 100 ml plasma with 200 ml buffer. Why?
The key is that the plasma that does not have a pre-dilution corresponds to 150%, not 100%. One way to look at it is that since 100 is 67% of 150, then the plasma must be diluted 67%, which corresponds to 100 ml plasma with 50 ml buffer (100 plasma / 150 total volume = 0.67). This process can be followed for all of the standard pre-dilutions.
Theoretical basis for calculation
The hydrolysis of the chromogenic peptide substrate by the proteolytic enzyme follows in general the Michaelis-Menten kinetics. This means that, if the substrate is present at a sufficiently high concentration or if a comparatively small fraction of the substrate is hydrolized, the rate of product (color) formation is proportional to the activity of the enzyme. The rate of pNA formation, i.e. the increase in absorbance per second, is measured photometrically at 405 nm. At this wavelength the extinction coefficient of pNA is
9600 mol -1 • l • cm -1
The enzymatic activity can be quantified in two ways:
- By comparing the activity of an enzyme with that of a standard preparation, which is defined in terms of a specified number of units set by an international or national authority or society (WHO, NIH etc.), or by the activity present in 1 ml of activated pooled normal plasma (Plasma Equivalent Unit = PEU). The standardization is performed by using at standard curve obtained with at least five different concentrations, each performed in duplicate. The standard material must be of the same kind and of the same quality as the sample which is to be measured. This may be still more important for a secondary or domestic standard.
- By measuring the amount (mol) of substrate split, or rather product formed per unit time (absolute activity).
One unit of enzymatic activity, katal (kat) is defined as the amount of activity that converts one mole of substrate per second under standardized conditions. Such conditions as type of substrate, substrate concentration, buffer, pH, ionic strength and temperature should be given along with unit.
Thus, 1 nkat gives a conversion rate of:
1 x 10-9 mol/sec = 60 x 10-9 mol/min
If the total (measuring) volume used is V (ml), the increase in concentration per minute caused by 1 nkat is
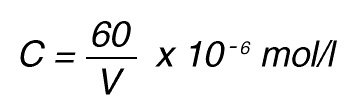
If the absorbance is measured at 405 nm, in a 1 cm cuvette the difference in extinction coefficient is
e = 9600 mol-1 • l
The increase in absorbance/min can then be calculated by using Lambert-Beer’s law:
A = e x C
Thus, 1 nkat gives:
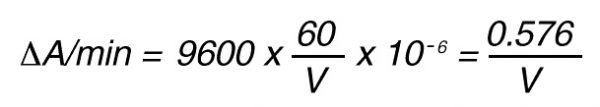
or
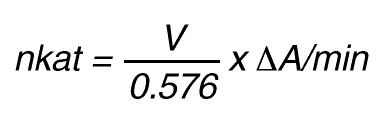
By using a sample volume v (ml):
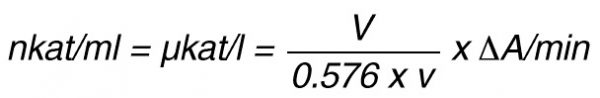
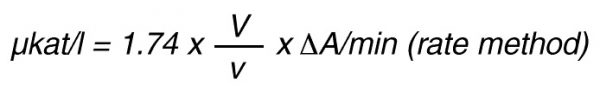
For the end-point method, the incubation time t (min) with substrate is taken into account by the following formula:
![]()
According to nomenclature, one unit (U) is the amount of enzyme activity that converts one mol of substrate per minute under standardized conditions. By using the above formulas the units are:
![]()
Products: S820316, S820324, S820332, S820340, S820852, S821090, S821413, S822254
There are a few different substrates that are hydrolyzed by plasmin. If I want to use as short incubation times as possible, and need a selective substrate for plasmin, which should I choose?
The substrates for plasmin include S-2251™, S-2302™, and S-2403™. While S-2251™ is a popular and suitable substrate for detection of plasmin, S-2403™ has a higher kcat/km value. It is a faster substrate, and incubation times can be shorter. S-2403™ is therefore the substrate of choice for this situation.
We know that low levels of factor VIII activity constitutes hemophilia A, but are there any clinical manifestations of elevated factor VIII levels? How can a researcher measure elevated FVIII levels?
There is evidence that a high level of factor VIII is a risk factor for venous thrombosis. There are two articles in Thrombosis and Haemostasis 2000; volume 83:
1.Kraaijenhagen et al. High plasma concentration of factor VIII:c is a major risk factor for venous thromboembolism (p. 5-9)
2.O’Donnell et al. Elevation of FVIII:c in venous thromboembolism is persistent and independent of the acute phase response (p. 10-13)
Both of these recent articles discuss the issue of elevated factor VIII levels as a risk factor for venous thrombosis.
An adaptation of Chromogenix Coamatic® FVIII has been developed to allow accurate determination of elevated factor VIII levels. The method is for research use only.
Products: K822585
What anticoagulants can be used with Multiplate® testing?
The instrument manufacturer has provided reference ranges for three anticoagulants; lithium heparin (typically a green top blood collection tube), 3.2 % citrate (typically a blue top blood collection tube), and an RUO hirudin blood collection tube (3 mL) available from the instrument manufacturer. With the hirudin whole blood, the hirudin concentration in whole blood is >15 ug/mL. Note that the ACD blood collection tube (typically a yellow top blood collection tube) is not suitable for platelet function testing as low pH inhibits platelet function. Other anticoagulants require user validation for their suitability.
What are anti-phospholipid antibodies and anti-phospholipid antibody cofactors?
Antiphospholipid antibodies are a heterogeneous group of auto-antibodies (IgG, IgM, and IgA) initially thought to be specific only to negatively charged phospholipids. It is now well recognized that many antiphospholipid antibodies are directed to phospholipid-protein complexes and /or to proteins in the absence of phospholipids.
Antiphospholipid antibody cofactors are plasma proteins mostly with function in coagulation and strong phospholipid binding activities. Cofactors increase the binding of antiphospholipid antibodies in vitro and may help the development of thrombosis in vivo.
What are autoantibodies?
Autoantibodies are antibodies to self components produced when immunologic tolerance is broken.
What are FVIII inhibitors, and how can I measure them?
FVIII inhibitors appear in about 25% of severe hemophilia A patients. They are most common in patients with inversions, large deletions and nonsense mutations. Inhibitor titer varies 1000-fold; high titer seriously impairs treatment efficacy. Inhibitors can be transient or persistent. and appear irrespective of which FVIII concentrate is selected. High titer patients are treated with high doses of human or porcine concentrate. View the FVIII product pages for kits and assays for FVIII Inhibitor measurements.
What are some features of the REAADS anti-Cardiolipin test? The b2GPI test?
The REAADS anti-Cardiolipin test kit and the b2GPI kit are reagent-complete kits. The anti-cardiolipin kit features specific determination of IgG, IgM, and IgA aCL antibodies. The kits are convenient, cost-effective ELISA procedures which give objective, accurate, and reproducible results with short incubations at room temperature.
Products: K023-001, K026-001, K037-001, K038-001, K039-001, 13525, 13529
What are some of the inhibitors to t-PA?
PAI-1, PAI-2, PAI-3, protease nexin, a2-macroglobulin, trypsin inhibitor, and C1 Inhibitor all inhibit tPA. Plasminogen activator inhibitor 1 (PAI-1) is the most efficient inhibitor of tPA in plasma. It is a serine protease inhibitor (serpin) that acts as a pseudo-substrate for its target protease, with which it forms an inactive complex. PAI-1 is synthesized by several cell types including endothelial cells and hepatocytes and is present in platelets, placenta, and serum. The normal concentration range of PAI-1 in plasma is 5-40 mg/l and the normal activity is 0-20 AU/ml.
PAI-2 is a serpin with a higher affinity for u-PA (urinary-type plasminogen activator, or urokinase) than for t-PA. It is often only detectable during pregnancy, specifically in the third trimester.
PAI-3 is also called Protein C inhibitor, and inhibits u-PA and thrombin, and is present in plasma and urine.
What are some phospholipid cofactors?
- beta2 Glycoprotein 1
- Prothrombin
- Protein C
- Protein S
- Annexin V
- Kininogen
What are the advantages of performing ELISA rather than molecular testing?
Molecular testing reflects the gene expression whereas ELISA assays detect the product, or actual effect, of this expression. Additionally, molecular assays tend to be time-consuming, costly and more sensitive to technique variables such that they require special laboratories with well-trained personnel.
Products: TE1037, TE1035, TE1034, TE1042, TE1040, TE1043, TE1046, TE1049, TE1039
What are the advantages of using mucus rather than blood or whole body homogenate?
Mucus collection is non-invasive and non-destructive. It can be easier to collect from smaller fish. Collectingmucus does not require sacrificing the fish which means repeat sampling allows for studying induction kineticsby individual. Repeat sampling also allows for pre-exposure testing, making it possible to assess handling(stress)-related impacts on VTG induction. Additionally, research indicates the vitellogenin protein may be morestable in mucus. In blood, the vitellogenin protein can be especially prone to proteolytic cleavage.
Products: TE1037, TE1035, TE1034, TE1042, TE1040, TE1043, TE1046, TE1049, TE1039
What are the different types of protein S deficiency?
The classification of PS deficiency is as follows:
Type 1: Low Protein S Ag Total, Low Protein S Ag Free, Low Protein S Activity
Type II: Normal PS Ag Total, Normal PS Ag Free, Low PS Activity
Type III: Normal PS Ag Total, Low PS Ag Free, Low PS Activity
Type II deficiency seems to be quite rare. Acquired PS is also possible, such as with liver disease, DIC, IBD, and APS. Some research suggests that free PS rather than total PS should be measured in the diagnosis of PS deficiency.
What are the latest recommendations for measuring potentcies of high purity factor VIII concentrates?
- A concentrate standard should be used in lieu of plasma standards.
- Sample and standard concentrates should be pre-diluted in Factor VIII deficient plasma
- All assay buffers should contain 1% (10 mg/ml) of high purity albumin, either bovine or human.
- The chromogenic method is the reference method.
What are the maintenance requirements?
On a daily basis when using the AR chip, the EDTA line/pump is purged with distilled water and stored in distilled water to prevent EDTA salt deposition.
On an annual basis, the instrument tubing and filter on the EDTA line/pump is replaced.
What are the materials needed for a PL assay or AR assay?
One PL chip package kit consists of 15 chips and there are 2 PL tests per chip. Each PL chip package requires PL sample reservoirs (30)/overcaps (30). The whole blood must be drawn in a hirudin vacuum collection tube (3 mL draw); these are packaged as 50 tubes.
One AR chip package kit consists of 15 chips and there is one AR test per chip. Each AR chip package requires AR sample reservoirs (15)/overcaps (15) and waste tanks (15). The whole blood must be drawn in a specific 3.2 % citrate vacutainer tube (4.5 mL draw, Terumo, 100 tubes). The assay also requires CTI/Ca solution (0.6 mL) and a solution of 25 mM Na EDTA which the user prepares.
Note that the sample reservoir differs for the PL and the AR chip. And that there are specific blood collection tubes to use for each test!
What are the reagent preparations before testing?
As the PL and the AR chip is stored at -20° C or colder, they should equilibrate at RT for 30 minutes before use. Heating the chip on the instrument preheater can accelerate the RT equilibration.
What are the test strips coated with?
What aspects must a scientist consider when choosing the best chromogenic substrate?
Synthetic substrates are very sensitive; they can detect very low enzyme activities and are often more sensitive than a corresponding natural substrate. On the other hand, they can be less selective, or, have less discrimination in their reactivities toward related enzymes compared to the natural substrate. There are steps a scientist can take to maximize sensitivity and specificity. If the specificity of the enzymatic activity to be measured is known then a substrate selectivity table which shows the cross-reactivity of the substrates with different enzymes, and the kinetic data, such as that provided in the Chromogenix catalog, can be helpful. If the specificity of the enzyme is unknown, a screening procedure can be applied. This involves comparing the rate of hydrolysis obtained with different substrates. The presence of contaminating enzymes must also be taken into account. To eliminate interference, an inhibitor can be introduced, the sample can be further diluted, or conditions can be found where the relative activities are optimized. For instance, S-2222™ is selective for FXa, but also for trypsin. If a researcher wants to measure FXa, s/he can add an inhibitor to trypsin, such as soybean trypsin inhibitor. Temperature, pH, buffers, and ionic strength can all affect the rate of hydrolysis and must be considered. Substrate concentration is also important, and a concentration of 2 x Km is usually appropriate. A good substrate has a low Km, meaning maximum reaction velocity is achieved at a low substrate concentration. In other words, the enzyme has a high affinity for the substrate. A high Kcat is also desired, which means the enzyme has a high turnover rate with the substrate (fast reaction).
Products: S820316, S820324, S820332, S820340, S820852, S821090, S821413, S822254
What can I use for calibrators & controls for Factor X?
Currently there are no calibrators or controls on the market that are assayed for chromogenic FX, just clotting FX. Most labs use normal plasma calibrators and controls. If you calibrate your assay using normal plasma, you can determine the % of normal FX in your patient sample. Calibration and control plasma is available from DiaPharma that are traceable to the International Standards supplied by NIBSC in accordance with WHO recommendations.
What CPT code should I use for Factor X?
You can bill to CPT Code 85260
Products: KDPGFX
What CPT codes should I use?
Search the AMA for CPT codes: https://catalog.ama-assn.org/Catalog/cpt/cpt_search.jsp
Current Procedural Terminology (CPT) Codes describe medical procedures performed by physicians and other health providers. The codes were developed by the Health Care Financing Administration (HCFA) (later renamed the Centers for Medicare & Medicaid Services) (CMS) to assist in the assignment of reimbursement amounts to providers by Medicare carriers. A growing number of managed care and other insurance companies, however, base their reimbursements on the values established by HCFA.
Since the early 1970s, HCFA has asked the American Medical Association (AMA) to work with physicians of every specialty to determine appropriate definitions for the codes and to try to determine accurate reimbursement amounts for each code. Two committees within CMS work on these issues: the CPT Committee, which updates the definitions of the codes, and the RUC (Relative Value Update Committee), which recommends reimbursement values to CMS based on data collected by medical societies on the going rate of services described in the codes.
Common Hemostasis CPT Codes
| Code | Description |
|---|---|
| 85307 | Activated Protein C (APC) resistance assay |
| 85410 | alpha-2 antiplasmin (fibrinolytic factors and inhibitors) |
| 0030T | Antiprothrombin (phospholipid cofactor) antibody, each Ig class |
| 86148 | Anti-phosphatidylserine (phospholipid) antibody |
| 85300 | antithrombin III, activity (clotting inhibitors or anticoagulants) |
| 85301 | antithrombin III, antigen assay (clotting inhibitors or anticoagulants) |
| 86146 | beta 2 Glycoprotein I antibody, each |
| 86141 | C-reactive protein; high sensitivity (hsCRP) |
| 86147 | cardiolipin (phospholipid) antibody, each Ig class |
| 85130 | Chromogenic substrate assay |
| 85378 | D-dimer; qualitative or semiquantitative (fibrin degradation products) |
| 85379 | D-dimer; quantitative (fibrin degradation products) |
| 85380 | D-dimer; ultrasensitive (eg, for evaluation for venous thromboembolism), qualitative or semiquantitative (fibrin degradation products) |
| 85293 | high molecular weight kininogen assay (Fitzgerald factor assay) (clotting) |
| 85210 | factor II, prothrombin, specific (clotting) |
| 85220 | factor V (AcG or proaccelerin), labile factor (clotting) |
| 85230 | factor VII (proconvertin, stable factor) (clotting) |
| 85240 | factor VIII (AHG), one stage (clotting) |
| 85244 | factor VIII related antigen (clotting) |
| 85245 | factor VIII, VW factor, ristocetin cofactor (clotting) |
| 85246 | factor VIII, VW factor antigen (clotting) |
| 85247 | factor VIII, von Willebrand factor, multimetric analysis (clotting) |
| 85250 | factor IX (PTC or Christmas) (clotting) |
| 85260 | factor X (Stuart-Prower) (clotting) |
| 85280 | factor XII (Hageman) (clotting) |
| 85290 | factor XIII (fibrin stabilizing) (clotting) |
| 85291 | factor XIII (fibrin stabilizing), screen solubility (clotting) |
| 85362 | Fibrin(ogen) degradation (split) products (FDP)(FSP); agglutination slide, semiquantitative |
| 85366 | Fibrin(ogen) degradation (split) products (FDP)(FSP); paracoagulation |
| 85370 | Fibrin(ogen) degradation (split) products (FDP)(FSP); quantitative |
| 85384 | Fibrinogen; activity |
| 85385 | Fibrinogen; antigen |
| 85520 | heparin assay |
| 85400 | plasmin (fibrinolytic factors and inhibitors) |
| 85415 | plasminogen activator (fibrinolytic factors and inhibitors) |
| 85420 | plasminogen, except antigenic assay (fibrinolytic factors and inhibitors) |
| 85421 | plasminogen, antigenic assay (fibrinolytic factors and inhibitors) |
| 85415 | plasminogen activator (fibrinolytic factors and inhibitors) |
| 85420 | plasminogen, except antigenic assay (fibrinolytic factors and inhibitors) |
| 85292 | prekallikrein assay (Fletcher factor assay) (clotting) |
| 85302 | protein C, antigen (clotting inhibitors or anticoagulants) |
| 85303 | protein C, activity (clotting inhibitors or anticoagulants) |
| 85305 | protein S, total (clotting inhibitors or anticoagulants) |
| 85306 | protein S, free (clotting inhibitors or anticoagulants) |
What dilution should be used to prepare samples?
The best sample dilution will vary based on type of sample (blood, homogenate or mucus), species and exposure to test compound(s). Refer to the package insert and/or the Vitellogenin ELISA Crossreactivity chart for suggested dilution ranges. During assay evaluation/validation, it is recommended that several dilutions, based on the recommended range, be tested initially using a subgroup of samples. The results from assaying the subgroup at several dilutions can then be used to determine the best dilution for the rest of your samples based on your experimental design.
Products: TE1037, TE1035, TE1034, TE1042, TE1040, TE1043, TE1046, TE1049, TE1039
What effect does variation of plasma levels of protein C have on the APC ratio?
What factors cause increased and decreased levels of t-PA and PAI?
There are numerous physiological factors that influence tPA and PAI antigen and activity level. For a complete list, please request the tPA monograph from DiaPharma. One interesting feature of the fibrinolytic system is the circadian variation in tPA and PAI-1level. Free tPA levels are lowest in the morning, increase during the day, and reach their peak activity level in the late afternoon. tPA and PAI-1 antigen are highest in the early morning and decrease during the day. This may help explain the high incidence of MI and stroke in the morning hours. Other factors that influence tPA and PAI-1 include alcohol, drugs, oral contraceptives, exercise, food, heparin administration, pregnancy, smoking, etc.
What if the reconstituted substrate has some precipitate or is cloudy?
The substrate solution is usually prepared with sterile water, but sometimes they may not dissolve properly. Sonication may help, or substrates with low solubility in water can be dissolved in DMSO, then diluted in water. The final DMSO concentration should preferably not exceed 10% in the reaction mixture. It should be noted that stability in DMSO is decreased, as it also is with alkaline buffers.
Products: S820316, S820324, S820332, S820340, S820852, S821090, S821413, S822254
What is a chromogenic substrate?
Enzymes are proteins that catalyze most of the chemical reactions that take place in the body. They make it possible for chemical reactions to occur at neutral pH and body temperature. The chemical compound upon which the enzyme exerts its catalytic activity is called a substrate.
Proteolytic enzymes act on their natural substrates, proteins and peptides by hydrolyzing one or more peptide bond(s).
This process is usually highly specific in the sense that only peptide bonds adjacent to certain amino acids are cleaved.
Chromogenic substrates are peptides that react with proteolytic enzymes under the formation of color. They are made synthetically and are designed to possess a selectivity similar to that of the natural substrate for the enzyme.
Attached to the peptide part of the chromogenic substrate is a chemical group which when released after the enzyme cleavage gives rise to color. The color change can be followed spectrophotometrically and is proportional to the proteolytic activity.
The chromogenic substrate technology was developed in the early 1970s, and has since then become a tool of substantial importance in basic research.
The majority of chromogenic substrate applications are found in various clinical fields. In particular they have been used to generate fundamental knowledge of the mechanisms regulating blood coagulation and fibrinolysis.
Furthermore, products based on chromogenic substrate technology have brought a new generation of diagnostics into the clinical laboratory.
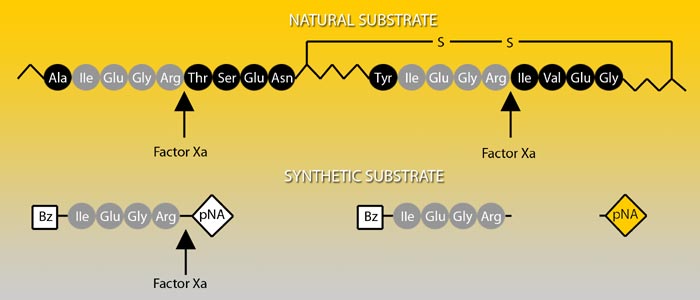
Prothrombin, the natural substrate of Factor Xa, is cleaved by Factor Xa at two positions, each proceeded by the same four amino acid sequence. Factor Xa activity can be determined by the chromogenic substrate S-2222™ which is composed of the same amino acids coupled to a chromophore.
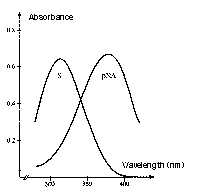
Absorption spectrum of a chromogenic pNA-containing substrate (S) and of pNA. The hydrolysis of a peptide-pNA bond in the chromogenic substrates results in the release of pNA which in turn changes color. Thus the change in absorbance (ΔA/min) is directly proportional to the enzymatic activity. The reaction is usually recorded at 405 nm.
Products: S820316, S820324, S820332, S820340, S820852, S821090, S821413, S822254
What is a chromogenic substrate composed of?
Chromogenic substrates are peptides that react with proteolytic enzymes under the formation of color. Chromogenic substrates are made up of a protecting group, amino acid residue(s), side-chain modification if applicable, and the chromophore. The stereochemistry of some substrates may be designated. For example, in the Chromogenix substrate S-2222™, the protecting group is a benzoyl group, the amino acid residue is Ile (isoleucine – a non-polar hydrophobic amino acid), the side chain modification is Glu(g-OR)- where R is 50% H (hydrogen) and CH3 (methyl group). The P2 and P1 amino acid residues are Gly and Arg, respectively, and the chromophore is pNA (para-nitroaniline).
Products: S820316, S820324, S820332, S820340, S820852, S821090, S821413, S822254
What is a peptide? What is the difference between a tripeptide and a tetrapeptide? How are amino acids linked to form peptides?
A peptide is the name assigned to short polymers of amino acids. They are classified by the number of amino acid units in the chain, called amino acid residues. Tripeptides have three amino acid residues while tetrapeptides have four. A polypeptide is formed when the chain of amino acid residues exceeds several dozen in length. A protein is a molecule composed of one or more polypeptide chains.
Proteins are unbranched polymers of amino acids linked head to tail from carboxyl group to amino group, through formation of covalent peptide bonds. The peptide backbone of a protein consists of the repeated sequence.
-N-Ca– C, where N represents the amide nitrogen, Ca represents the a-carbon atom of an amino acid in the polymer chain, and the final C is the carbonyl carbon of the amino acid. This C is in turn linked to the amide N of the next amino acid, and so on down the line. The unbranched polypeptide chain has two ends, an amino-terminal or N-terminal end and a carboxyl-terminal or C-terminal end.
Products: S820316, S820324, S820332, S820340, S820852, S821090, S821413, S822254
What is a proteolytic enzyme?
In the living organism, proteolytic enzymes (proteases) are produced to degrade and modify proteins. A main task for proteolytic enzymes is to degrade proteins into peptides or amino acids to be used either as an energy source or as building blocks for resynthesis of proteins. Furthermore, proteolytic enzymes modify cellular environments and facilitate cell migration in connection with wound repair and cancer, ovulation and implantation of the fertilized egg, embryonic morphogenesis, and involution of mammary glands after lactation.
Another important function of the proteases is their role as regulators in processes such as inflammation, infection and blood clotting. Most proteolytic enzymes are highly specific for their substrates. The classification of proteases, however, is not based on their choice of substrate but on their mechanism of action.
Four different groups of proteolytic enzymes, named after the active site amino acid residue responsible for the catalytic activity, are generally distinguished: the aspartic proteases (e.g. pepsin), the cystein proteases (e.g. cathepsin B and cathepsin H), the serine proteases (e.g. trypsin, thrombin and plasmin) and metalloproteases (e.g. collagenases and gelatinases).Although the members of each group of proteolytic enzymes may have very diverse biological functions, amino acid analysis often shows a high degree of structural similarity between them. Detailed knowledge of the structure and mechanism of action of one enzyme can in many cases reveal an understanding of the structure and functions of other enzymes within the same group.
Classes of Proteases
| Name | active site |
|---|---|
| serine proteases | Ser His Asp* |
| cystein proteases | Cys His Asp* |
| aspartic proteases | Asp Asp |
| metallo proteases | His His Zn2+ |
*Asp not always present
What is a serine protease?
The most extensively studied group of proteolytic enzymes comprises the serine proteases. As indicated by the name each member of this group have a reactive seryl amino acid residue in its active site.
The serine proteases are divided into two families: the trypsins and the subtilisins.
The trypsin family is the largest and contains, among others, trypsin and chymotrypsin, elastase, mast cell tryptase, and many of the factors regulating blood coagulation and fibrinolysis.
The trypsin type of enzymes have a highly similar amino acid content. They are found in vertebrates and other animals, as well as in fungi and procaryotic cells. In contrast, the subtilisins are only found in bacteria. Members of the trypsin family are classified according to the type of amino acid that occurs at the preferred cleavage site.
Elastase and chymotrypsin cleave after hydrophobic and aromatic amino acids, while other trypsin-like proteases cleave only at the C-terminal side of the basic amino acids arginine or lysine. The amino acid sequence and thus also the three-dimensional structure differ completely between the trypsins and the subtilisins. The catalytically active domains of trypsin and subtilisin have therefore most probably evolved independently, converging from two different genes.
However, since the three amino acids of functional importance at the active sites, serine (Ser), aspartic acid (Asp) and histidine (His), are arranged in the same geometrical relationship in all members of the two families the proteolytic mechanisms are very similar.
This fact may lead to the suggestion that the arrangement of the three catalytically active amino acids at the active site is very efficient for hydrolysis of peptide bonds. Mammalian serine proteases are usually synthesized as inactive proenzymes, zymogens, consisting of a single peptide chain. Activation occurs when the zymogen is cleaved at one or several specific sites. Most commonly such cleavage is accomplished by the action of another protease. Most serine proteases contain two functionally distinct parts.
The region where the catalytically active amino acids are found is very similar in trypsin and chymotrypsin as well as in the serine proteases involved in blood coagulation. The other region is located in the exterior parts of the enzyme. This region is of considerable size in the serine proteases regulating blood coagulation and fibrinolysis and four main types of structures can be distinguished: kringle domains, growth factor domains, vitamin K dependent carboxylated calcium binding domains, and domains homologous to the finger structure of fibronectin.
All four domain types are not present in all groups of serine proteases.
In the living organism, proteolytic enzymes (proteases) are produced to degrade and modify proteins. A main task for proteolytic enzymes is to degrade proteins into peptides or amino acids to be used either as an energy source or as building blocks for resynthesis of proteins. Furthermore, proteolytic enzymes modify cellular environments and facilitate cell migration in connection with wound repair and cancer, ovulation and implantation of the fertilized egg, embryonic morphogenesis, and involution of mammary glands after lactation.
Another important function of the proteases is their role as regulators in processes such as inflammation, infection and blood clotting. Most proteolytic enzymes are highly specific for their substrates. The classification of proteases, however, is not based on their choice of substrate but on their mechanism of action.
Four different groups of proteolytic enzymes, named after the active site amino acid residue responsible for the catalytic activity, are generally distinguished: the aspartic proteases (e.g. pepsin), the cystein proteases (e.g. cathepsin B and cathepsin H), the serine proteases (e.g. trypsin, thrombin and plasmin) and metalloproteases (e.g. collagenases and gelatinases). Although the members of each group of proteolytic enzymes may have very diverse biological functions, amino acid analysis often shows a high degree of structural similarity between them. Detailed knowledge of the structure and mechanism of action of one enzyme can in many cases reveal an understanding of the structure and functions of other enzymes within the same group.
What is enzyme specificity and substrate selectivity?
Specificity is a property of the enzyme and describes how restrictive the enzyme is in its choice of substrate; a completely specific enzyme would have only one substrate.
The specificity of the serine proteases is usually not very high since they have similar active sites and act through the same proteolytic mechanism.
Consequently, a single serine protease may act on various substrates although at different rates. How the substrate fits the active site of the enzyme is of crucial importance to the outcome of the enzyme-substrate reaction. The bond to be cleaved must have a specific orientation relative to the amino acid side chains of the catalytic triad. The most important factor governing the fit of a substrate for an enzyme is the amino acid sequence around the bond to be cleaved.
Trypsin cleaves amides and esters of the basic amino acids arginine and lysine. Thrombin has a similar preference, but is more specific for arginine than for lysine.
Selectivity is a property of the substrate and indicates the degree to which the substrate is bound to and cleaved by different enzymes. The best measure for selectivity is given by the ratio kcat/Km. Synthetic substrates are considerably smaller than the natural substrates and can usually be cleaved by more than one enzyme, i. e. synthetic substrates are not completely selective. The explanation for this is that large substrates such as fibrinogen not only interact with the active site but also with exterior domains of the enzyme. Such interactions allow substrates to discriminate between different serine proteases and fibrinogen thus becomes highly selective for thrombin.
Selectivity Tables
The selectivity data of the table have been compiled to permit the investigator to understand how a contaminating enzyme would influence the enzyme-substrate reaction under study. Another way of expressing this is to say that the table shows the relative reactivities of two or more enzymes on one particular substrate. The table should be read horizontally. Each row represents the reactivity of a substrate designated for use with a particular enzyme, indicated to the left, relative to other relevant enzymes.
Example: The set of data in the top row shows the relative reactivity of the thrombin substrate S-2238™ with various enzymes. All the experiments were performed using the same buffer, i.e. the one most appropriate for the reaction between thrombin and chromogenic substrate S-2238™. In addition, the substrate concentration was always the same, or 2 x Km for the reaction of chromogenic substrate S-2238™ with thrombin. The concentrations of the different enzymes are given in Table 2 and are related to the plasma concentration of the corresponding zymogen. The reactivity of chromogenic substrate S-2238™ with thrombin, measured as the time-dependent increase in absorbance (ΔA/min), is given the value 100% (the actual value of ΔA/min is given in brackets). The reactivities of chromogenic substrate S-2238™ with the enzymes FXa, FXIa, APC, plasmin, single chain t-PA, plasma kallikrein, and C1s have then been related to the reactivity of chromogenic substrate S-2238™ with thrombin, and proved to be 5, 5, 40, 5, 5, 60, and 2%, respectively.
What is heparin-induced thrombocytopenia (HIT)? When HIT is suspected in a patient treated with UF heparin, should LMW heparin therapy replace UF therapy?
HIT, defined by the presence of heparin-dependent IgG antibodies, is characterized by a decrease in platelet count shortly after starting heparin, which resolves after stopping heparin and is not due to any other apparent cause. Mild HIT occurring within 2-3 days is due to a direct effect of heparin on platelets and is not immune-mediated nor associated with thrombosis. Severe thrombocytopenia, usually occurring a few days later, is associated with both arterial and venous thrombosis and is immune-mediated. In most cases, it results from antibody formation to heparin-platelet factor 4 (PF4) complexes, but in about 10% of cases heparin appears to bind to pre-existing antibodies. Although LMW heparin seems to cause fewer incidences of HIT, it should not replace UF heparin therapy if HIT is already suspected. It may exhibit in vitro and in vivo cross-reactivity with UF heparin-dependent antibodies. Therefore, when HIT is suspected, other anticoagulant options must be explored.
What is the 53 nkat vial of thrombin equivalent to in NIH units?
What is the advantage of an assay for anti-phospholipid antibody cofactors over an assay for the antibody itself?
Binding of b2GPI to the microwell surface in an ELISA assay may produce a neoepitope similar to that when combined with a phospholipid; assay results with this system show a good correlation with the APS. The serologic detection of anti-b2GPI antibodies provides enhanced clinical sensitivity for thrombosis.
What is the advantage of using the Chromogenix Coamatic® FVIII over a clotting test?
While clotting assays are cheap, rapid, and simple to perform, they are sensitive to pre-activation, show interferences in the assays, and can give overestimation of FVIII concentrates. They also require a considerable amount of FVIII deficient plasma.
Coamatic® FVIII is also rapid and simple, and has additional benefits. It is suited for a wide range of automated instrument applications, there is no heparin influence, and it is highly sensitive, precise and accurate. The low measurement range provides a reliable tool for the classification of bleeders.
Products: K822585
What is the advantage of using the Peviva M30 CytoDeath™ ELISA relative to other methods to quantify apoptotic cell death?
The M30 CytoDeath™ ELISA measures a stable and abundant caspase cleavage product. It offers a very high sensitivity and makes it the method of choice for cost effective drug screening at a functional level, to establish dose-response curves and to establish time course kinetics in cellular systems in therapeutic relevant areas as oncology.
Products: P10900
What is the calibrator material?
The calibrator material is biologic and not recombinant. It is prepared from serum that has been run through a column and separated via affinity chromatography. It is extensively validated and characterized using immunohistochemical staining, Western blot and SDS PAGE technology.
Products: TE1037, TE1035, TE1034, TE1042, TE1040, TE1043, TE1046, TE1049, TE1039
What is the Chromogenix Coamatic® Factor VIII measurement principle?
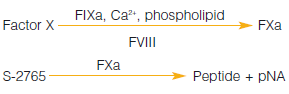
Products: K822585
What is the Chromogenix Coamatic® Heparin measurement principle?
![]()
Products: K823393
What is the Chromogenix Coatest® Heparin measurement principle?
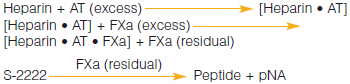
Products: K255539
What is the clinical significance of Protein S?
Although the exact role of PS in vivo in the protein C anticoagulant pathway has not been clarified, there is no doubt that PS is an important anticoagulant protein and that PS deficiency is primarily associated with venous thromboembolism. The reported prevalence of PS deficiency in thrombosis patients varies between 1.5-7%, the difference being due to methods used and to the selection of patients. Clinical symptoms in patients affected with PS deficiency are very similar to those with protein C deficiency (DVT). Superficial thrombosis seems to be more common than for antithrombin deficient patients.
What is the clinical significance when samples test positive for the anti-cardiolipin antibodies and are negative for anti-b2GPI?
The aCL assay can detect antibodies of differing specificities. This includes antibodies specific for the cardiolipin molecule itself, and antibodies that are directed against either a cofactor molecule such as b2GPI, or a special binding site created by the interaction of a cofactor with CL. Antibodies directed against CL may be associated with infectious disease, or may be specific for a different cofactor, such as prothrombin. The clinical significance of these antibodies must be assessed in conjunction with the patient’s symptoms, clinical history, and other laboratory findings. Follow-up testing of these patients is recommended in 3-6 months to confirm reactivity. Only b2GPI cofactor dependent antibodies react in the anti-b2GPI assay; these antibodies show a higher correlation with thrombosis and are more specific for the antiphospholipid syndrome.
Products: K023-001, K026-001, K037-001, K038-001, K039-001, 13525, 13529
What is the composition of mucus extraction buffer and sample dilution buffer?
What is the composition of the matrix solution and what is its purpose in the assay?
What is the conversion factor for Antithrombin from IU to mg?
What is the DiaPharma Factor X kit?
The DiaPharma Factor X Kit is an in vitro diagnostic test kit for the quantitative determination of Factor X activity in human citrated plasma. Factor X activity is useful for monitoring patients on oral anticoagulant therapy (warfarin) where baseline INR values are prolonged and INR results are not reliable, such as in OAC patients with lupus inhibitors. The DiaPharma Factor X Kit is also useful for screening for factor X deficiencies.
Products: KDPGFX
What is the difference and advantage of the Peviva M30 CytoDeath™ ELISA compared to assays that measure e.g. caspase-3 activity?
The caspase-3 assay measures enzyme activity (with specificity problems associated with “peptide caspase”-substrates), while the M30 CytoDeath™ ELISA measures the accumulation of a caspase cleavage product. Due to the accumulation of the analyte (K18Asp396-NE) over time, M30 CytoDeath™ ELISA is more sensitive and does not require consideration of the optimal analytical (often transient) window. This makes this assay convenient and economical when a large number of samples are analyzed. A limitation of the M30 CytoDeath™ ELISA is that it will only work with epithelial cells.
Products: P10900
What is the difference between an ELISA test and a chromogenic test? Which of our kits uses which technology?
An ELISA (enzyme linked immunosorbent assay) is an antigenic test which gives a quantitative measurement of the analyte antigen. It will measure active as well as latent forms of the analyte. In contrast, a chromogenic assay measures functional activity. A patient could have normal levels of an analyte, such as PAI-1 antigen, but there could be some mutation on the molecule leading to a decrease in analyte activity. An ELISA would therefore detect nothing abnormal, whereas a chromogenic assay would.
What is the difference between Chromogenix Coamatic® FVIII, Coatest® SP FVIII, and Coatest® SP FVIII?
All of these kits measure the same thing — factor VIII activity in plasma and potency estimation of FVIII concentrates. Coamatic® FVIII includes the FXa substrate S-2765™, and the factor reagent FIXa + FX and thrombin colyophilized with CaCl2 and phospholipid. The Coatest® SP FVIII includes the FXa substrate S-2765™, and the FIXa + FX reagent, CaCl2, and phospholipid are all separate components in the kit. Incubation times are a little longer, and the reagent prep is less simple. Coatest® SP4 FVIII is exactly the same as Coatest® SP FVIII, but the FIXa + FX reagent is divided into 4 vials of 1.8 U (reconstituted with 3 ml) instead of 1 vial of 6 U (reconstituted with 10 ml).
What is the difference between single and two-chain tPA?
What is the function of antithrombin and what is its interaction with thrombin and heparin?
Antithrombin is the most important natural inhibitor of the coagulation cascade, accounting for approximately 80% of the thrombin inhibitory activity in plasma. By inhibiting the coagulation proteases, especially thrombin, FXa, and FIXa, AT prevents uncontrolled coagulation and thrombosis. Inhibition of antithrombin involves the formation of a stable 1:1 complex between the active domain of the serine protease such as thrombin, and the reactive site of antithrombin, which proteases initially recognize as a substrate. During the cleavage of the reactive site bond in antithrombin, a conformational change occurs in the inhibitor that traps the protease.
Slow protease-antithrombin interactions are enhanced dramatically in the presence of certain sulfated polysaccharides like heparan sulfate. Heparin is a commercial preparation of heparan sulfate, and binds antithrombin and thrombin, thereby catalyzing the thrombin-AT reaction. Binding to antithrombin induces a conformational change in AT that facilitates its reaction with thrombin. Thrombin binds to heparin in a non-specific manner and slides along the chain until it encounters the bound AT. The affinity of heparin to the thrombin-AT (TAT) complex is much lower than to free AT. Heparin will therefore dissociate from the TAT complex, which is rapidly removed from the blood circulation by the liver.
What is the function of von Willebrand Factor? Briefly describe von Willebrand disease.
von Willebrand factor (vWf) is a glue-like adhesive protein that is responsible for the adhesion of platelets to damaged vascular endothelium. It also carries and protects factor VIII. von Willebrand disease is a hereditary bleeding disorder caused by moderate-to-severe factor VIII deficiency and low-levels of factor VIII-related antigen (substances necessary for blood clotting). Additionally, there is insufficient von Willebrand factor which also helps blood clot. The von Willebrand factor helps platelets to stick to the blood vessel wall and to each other, which is necessary for normal blood clotting.
Products: 5344101, 5450301, 5450311, 5450321, K034-001, K10826, K822585, K824086, K824094, 5450201, 800070, 13.2.095.0096
What is the importance of t-PA and PAI in clinical research?
What is the intended use of the Peviva M30 Apoptosense® assay?
The M30 Apoptosense® ELISA is intended for the quantitative detection of the apoptosis-associated M30 neo-epitope (K18-Asp396-NE) in blood samples.
Products: P10011
What is the intended use of the Peviva M30 CytoDeath™ ELISA?
The M30 CytoDeath™ ELISA is intended for the quantitative detection of the apoptosis-associated M30 neo-epitope (K18Asp396-NE) in cell extracts or cell culture supernatants. Potential applications include HTS screening for apoptotic drug candidates, performing time course kinetics and measuring dose responses after exposure to pro-apoptotic agents.
Products: P10900
What is the interaction between Protein S and APC?
Only the free, native form of PS binds to APC and functions as a cofactor. PS has the highest affinity for negatively charged phospholipids of all the vitamin K-dependent proteins and has been shown to increase by approximately 10-fold, the affinity of APC for membranes or vesicles containing such phospholipids. This may be of physiological importance since APC degrades preferentially membrane-bound FVa and FVIIIa, but not the circulating, inactivated co-factors. In addition to increasing the affinity of APC to membranes, PS also enhances the cleavage of FVa by APC and works in concert with FV to increase the ability of APC to inactivate FVIIIa.
What is the measurement principle behind the FVIII chromogenic kits?
First, factor VIII circulates in the plasma bound to von Willebrand factor (vWf). Thrombin cleaves and activates factor VIII and releases vWf. The vWf is then free to bind to ruptured endothelial cell surfaces where it activates platelet aggregation. The released FVIIIa acts as a cofactor of factor IXa to generate factor Xa. In the presence of Ca2+ and phospholipids, FX is activated to FXa by FIXa (look at the coagulation cascade for a better understanding). Since FVIIIa is a cofactor to FIXa, it greatly stimulates the reaction. By using optimal amounts of Ca2+, phospholipid, and FIXa, and an excess of FX, the rate of activation of FX is linearly related to the amount of FVIII. FXa hydrolyses the chromogenic substrate S-2765™ which releases the chromophoric group pNA. The color is read at 405 nm, and generated FXa and thus the intensity of color, is proportional to the FVIII activity in the sample.
What is the mode of action for the Peviva M30 CytoDeath™ ELISA?
M30 CytoDeath™ ELISA is a sandwich ELISA based on two mouse monoclonal antibodies, M30 and M6. The M30 antibody recognizes a neo-epitope in the C-terminal domain of keratin 18 (amino acids 387-396: K18Asp396-NE), exposed after cleavage by multiple caspases during apoptosis. The M6 antibody is used as capture antibody in the ELISA. Caspases that are capable of cleaving K18 to generate this M30 neo-epitope include in addition to caspase-3, also caspase-6, -7, and -9.
Products: P10900
What is the mode of action of the Peviva M30 Apoptosense® ELISA?
M30 Apoptosense® assay is a sandwich ELISA based on two mouse monoclonal antibodies, M30 and M5. The M30 antibody recognizes a neo-epitope in the C-terminal domain of keratin 18 (amino acids 387-396, K18Asp396-NE), exposed after cleavage by multiple caspases during apoptosis. The M5 antibody is used as capture antibody in the ELISA.
Products: P10011
What is the normal baseline range for the APTT reaction? What types of conditions might cause a patient to give abnormal aPT time, and what can be done about it?
The normal range is approximately 25-40 seconds, although it will vary slightly depending on instrument. Although the 1:4 dilution with V deficient plasma strongly decreases interferences from oral anticoagulants and heparin therapy, it cannot be excluded that analysis from patients with high inhibitor activity such as those with phospholipid antibodies (i.e. lupus anticoagulant) may give an abnormal APTT. Increasing the dilution factor to 1:9 or 1:19 may correct the results. According to a study by Nowak et al, a 1:10 dilution may also be useful in children less than one year old. This is due to the special properties of the neonatal hemostatic system, such as low vitamin-K dependent coagulation factors and physiological prolongation of the PT and APTT.
What is the proteolytic mechanism of serine proteases?
The bond-cleaving reaction exerted by a serine protease on its substrate is the result of an interaction between the substrate and the charge relay network of the enzyme. This network, which is present in the active site of all serine proteases, is known as the catalytic triad.
It is built up from the side-chains of three specific amino acids (the hydroxy group of serine, the imidazole group of histidine and the carboxylic acid group of aspartic acid) that interact with each other through an array of hydrogen bonds.
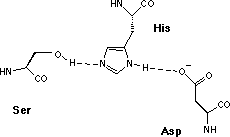
Charge relay network of serine proteases.
The proteolytic action of a serine protease on its substrate comprises several steps starting with the formation of a non-covalent complex between the enzyme and the substrate. A nucleophilic attack by the serine hydroxyl group on the amide carbonyl carbon atom in the substrate results in cleavage of the amide bond and the formation of an acyl-enzyme intermediate.
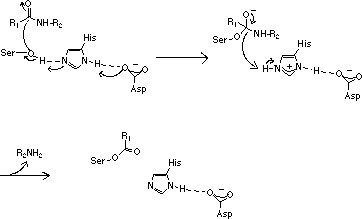
Formation of an acyl-enzyme intermediate.
The acyl-enzyme ester bond is then hydrolyzed in the rate limiting step and the enzyme is now free to catalyze the cleavage of another substrate molecule.
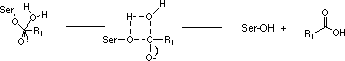
Hydrolysis of the acyl-enzyme intermediate.
What is the role of Beta 2 Glycoprotein 1 (B2GP1) in anti-phospholipid antibody binding?
b2GPI, also called apolipoprotein H, is an antiphospholipid protein cofactor with natural anticoagulant properties and an affinity for negatively-charged phospholipids. Antibodies directed against b2GPI have been shown to be specific markers for thrombosis in individuals with systemic lupus erythematosus (SLE) and lupus-like disorders (APS). Most autoimmune anti-phospholipid antibodies require the serum cofactor b2GPI for optimal binding. It has been shown that many anti-phospholipid antibodies may react to a neoepitope formed on the b2GPI molecule by the interaction between the phospholipid and b2GPI. The physiological role of b2GPI may additionally be to participate in apoptosis.
What is the species specificity of the Peviva M30 Apoptosense® assay?
The M30 antibody recognizes caspase-cleaved keratin 18 (K18-Asp396-NE M30) from human and monkey origin. K18-Asp396-NE from mouse, rat, dog or xenopus cells is not effectively detected.
Products: P10011
What is the structure and function of plasminogen?
Plasminogen is the inactive precursor of plasmin, a potent serine protease involved in the dissolution of fibrin clots in blood. Human plasminogen is a single-chain glycoprotein with a molecular mass of 92 kDa. The N-terminal portion of the plasminogen molecule consists of five kringle domains with the capacity to bind to fibrin. The kringles control the ability of plg to adopt different conformations. The protease domain resembles that of the other serine proteases and contains the active site His, Asp, Ser. The region also contains Ala, which appears to be essential for the normal function of plasminogen, since mutation from Ala601 to Thr601 (alanine to threonine point mutation) leads to an increased risk of thrombosis.
What is the therapeutic range for Factor X?
Each lab must determine its own reference range. Studies have shown that normal FX levels are about 70 – 130%, and therapeutic range is about 20 – 40%. A list of reference ranges found in the literature and at various reference laboratories can be obtained from DiaPharma.
Products: KDPGFX
What is the utility of the DiaPharma Factor X kit? Am I expected to use this on ALL my Coumadin patients?
No, only for those where the INR is unreliable and unstable. For example, it is useful in patients with lupus anticoagulant where the LA falsely prolongs the PT, therefore giving unreliable INRs. Also, Argatroban has been shown to prolong the PT, so patients being bridged from Argatroban to warfarin can be monitored using the DiaPharma Factor X kit.
Products: KDPGFX
What kind of samples can be analyzed by the Peviva M30 CytoDeath™ELISA?
As the caspase-cleaved product is released from cells, the antigen is present in the cell culture supernatant as well as in cytosolic cell extracts. Both types of samples can therefore be analyzed individually, or as a combined sample (by direct addition of detergent to the cell culture).
Products: P10900
What should I do for standards and controls for Factor VIII?
DiaPharma has calibrators, normal and abnormal controls available. Clinicians can also use their own pooled normal plasma; however, it must be calibrated against an international standard to be sure what concentration the plasma has. If the normal plasma does not contain exactly 1 IU/ml ( which corresponds to 100%) factor VIII, the values of the standards must be recalculated in order to obtain a correct factor potency assignment. Reference plasma is available from the National Institute for Biological Standards and Controls (NBSB). The calibration should be performed by running complete standard curves on two or more independent occasions. Industrial manufacturers, in contrast, use for their standard Mega-2, which is a compilation of factor VIII concentrate.
What substrate research methods are available?
- Chromogenix Chromogenic Substrate S-2222 Trypsin
- Chromogenix Chromogenic Substrate S-2222 Tissue Factor Pathway Inhibitor, type – I (TFPI)
- Chromogenix Chromogenic Substrate S-2238 Antithrombin
- Chromogenix Chromogenic Substrate S-2238 Antithrombin (anti-FIIa)
- Chromogenix Chromogenic Substrate S-2288 Proteolytic activity
- Chromogenix Chromogenic Substrate S-2238 Thrombin
- Chromogenix Chromogenic Substrate S-2228 Tissue Plasminogen Activator (t-PA)
- Chromogenix Chromogenic Substrate S-2251 Plasmin
- Chromogenix Chromogenic Substrate S-2302 Kallikrein
- Chromogenix Chromogenic Substrate S-2302 Kallikrein Inhibitor
- Chromogenix Chromogenic Substrate S-2302 Prekallikrein
- Chromogenix Chromogenic Substrate S-2366 Hirudin
- Chromogenix Chromogenic Substrate S-2765 Factor X
- Diapharma Chromogenic Substrate CS GK (substitute for discontinued Chromogenix S-2266) Kallikrein in urine
- Diapharma Chromogenic Substrate CS UK (substitute for discontinued Chromogenix S-2444) Urokinase
- Diapharma Chromogenic Substrate CS PSA (KLK3) (substitute for discontinued Chromogenix S-2586) Chymotrypsin
Historically, the development of a new chromogenic substrate for a specific protease has always been accompanied by the release of method sheets where the application and the methodology for a particular use were described in detail. Some methods are simple chromogenic assays where a buffer and the substrate are the only reagents to be used (i.e. proteolytic activity). Other methods instead require the use of additional compounds, which are or have been commercially available from Chromogenix or elsewhere, and consist of more reaction steps. These protocols were validated in laboratories according to the equipment and the reagents available at the time. In several cases they have been adopted in research, quality control, or routine laboratories and some of them later became Chromogenix kits now present in our product range.
During the last 20 years, the so-called Method Sheets have been taken as the starting point by several scientists, for the development of assays for particular applications, or studies. In some cases the experimental conditions have been changed according to the particular need of the investigation being done. These methods are now presented in a different form: “Research Methods”. The intention here is to provide assay protocols that are not available as kits, but complementary to our product range. In the following list, you can find the methods developed by Chromogenix for several assays. They have to be considered as general guidelines or basic tools for the development of your own assays, some of them requiring a validation within your laboratory with respect to the reagents and equipment used.
For each method, you can find an updated Bibliography with references, where the method has been used, like as originally described or with modifications. This information should facilitate and accelerate the development of the best test protocol. If you do not have that specific journal issue in your laboratory, you can visit MEDLINE. You can search for particular articles, print the abstract and order the original copy through LOANSOME DOC (and receive the document through your local library). At the same time, on our web site you have the possibility to be updated on the new products from Chromogenix.
Products: S820316, S820324, S820332, S820340, S820852, S821090, S821413, S822254
What type of Fluorescent Reader do I need?
While the manufacturer currently recommends using the Ceveron® alpha TGA analyzer, alternative fluorescent readers can be used. Excel data evaluation templates have been created for the BioTek® FLx800™, Spectramax M5, BMG FLUOstar, Perkin Elmer Wallac Viktor, TECAN, and Thermo Fluoroskan.
Products: 5006010
When exogenous AT is desired, how should it be added in the Chromogenix Coamatic® Heparin test?
Add to the assay an equal volume of 1 IU/ml AT as the plasma volume. AT, 10 IU, can be bought from DiaPharma. Whenever a sample is tested with exogenous AT, it should be measured against a standard curve also run with exogenous AT.
Products: K823393
Where is the TGA evaluation software?
The manufacturer refers to the Microsoft Excel data analysis templates as evaluation software. These are templates created by Technoclone that can be used for data analysis with a number of fluoresecent readers. The excel templates are available at www.technoclone.com or you can request one at info@diapharma.com.
Do not analyze your data with standard Microsoft Excel.
Products: 5006010
Which assay should I run Chromogenix Coatest® APC Resistance or Coatest® APC Resistance V?
During the last years recent publication have been showing that the APC Resistance phenotype is a risk factor for venous thrombosis, irrespective of the FV Leiden mutation, and hence this illustrates that the Coatest® APC Resistance and the Coatest® APC Resistance V kits measure different entities. For this reason, both Coatest® APC Resistance and the Coatest® APC Resistance V kits are included in the thrombophilia screening panel in some important laboratories. A recent and interesting publication on inherited thrombophilia was done by Uri Seligsohn and Aharon Lubetsky. In this paper, both APCR and APCR V are suggested as part of six high priority tests. The diagnosis for factor V Leiden should be confirmed by genetic test, in order to decide whether the family members should be examined.
Which of DiaPharma's kits can measure LMW heparin?
All of the heparin kits from DiaPharma can. Obviously, when measuring LMW heparin samples, LMW heparin standard, like Fragmin, must be used unless a specific universal calibrator is used. DiaPharma currently has calibrators available for LMW heparin and UF heparin.
Which substrate is best suited for measuring two-chain tPA, and why?
S-2765™, S-2366™, and S-2288™, S-2403™ are suitable for single-chain tPA, and S-2251™ was used in the discontinued Chromogenix Coaset® tPA kit, but there are no substrates specifically for two-chain tPA. A paper by Verheijen et al. (Thromb Res, 1985; 39: 281 – 288), however, describes a method comparing the direct amidolytic activity of tPA on S-2366™ and the plasminogen activating activity. Also, the substrate S-2288™ is suitable for measuring double-chain tPA because it has a slightly higher sensitivity than S-2366™. S-2288™ should be used with purified systems, though, since this substrate is sensitive for several proteases.
Products: S820332, S820852, S821090, S821413, S822254, TC12007, TC12080, TC16000
Who can benefit from using chromogenic factor VIII assays?
Clinicians or researchers interested in determining FVIII levels in patients with hemophilia A or with elevated factor VIII levels, which may be associated with thrombotic risk. Also, manufacturers interested in determining factor VIII potencies. The chromogenic FVIII activity method has been selected as the reference method by the EP for potency estimation of VIII concentrates, and it is a valuable tool for diagnosis an monitoring of hemophilia and for thrombophilia screening.
Why are the reactions measured at 405 nm?

The absorption intensity is expressed by the Beer-Lambert law, A = e x c x l, where A is absorbance, c is molar concentration, l is the path length of sample cell (usually 1 cm), and e is the extinction or molar absorptivity coefficient. The absorption spectrum of the substrate versus the chromophore, pNA (the chromogenic leaving group) is the reason for reading the reactions at 405 nm. The absorbance maximum of the unhydrolyzed, intact substrate is 316 nm and 380 nm for pNA. Although the difference between substrate and product is maximal at 385 nm, at 405 nm, there is less background reading, and the absorbance of the substrate is still less than 1% of that of an equimolar amount of pNA.
Products: S820316, S820324, S820332, S820340, S820852, S821090, S821413, S822254
Why are there 2 M65 ELISAs?
The M65® ELISA was introduced in 2004 and intended for the determination of total cell death of epithelial cells. A combined use of M65® ELISA and M30 Apoptosense® ELISA facilitates the determination of cell death mode (apoptosis or necrosis).
The M65 EpiDeath® ELISA is based on the same monoclonal antibodies as the M65® ELISA, except mAb M5 is used as the capture antibody (mAb M6 is used as capture antibody in the M65® ELISA). Reaction buffers have also been slightly modified This CK18 kit is manufactured differently and thus measures the (complex bound) CK18 epitopes differently than the M65® ELISA, which leads to a better discrimination between low or normal and high levels. It has a broad range as well.
The M65 EpiDeath ELISA is recommended whenever a customer wants to identify cell death with high sensitivity and specificity. The M65 ELISA is the better choice for customers intending to determine the cell death mode when using in combination with the M30 Apoptosense ELISA, especially when using a mathematical approach to describe the distribution between apoptosis and necrosis.
Why are there different anticoagulants to use with each chip?
The PL test is designed to measure primary hemostasis. The coagulation protein cascade is inhibited by adding hirudin (a direct thrombin inhibitor) to the whole blood. Platelets are activated by binding to the collagen-coated chip and the activated platelets aggregate as a white cellular clot.
The AR test is designed to measure cumulatively primary and secondary hemostasis. Platelets aggregate as the AR test is a collagen-coated chip and it is also coated with tissue thromboplastin. As the whole blood is collected in 3.2% citrate, calcium must be added at the start of the assay to allow secondary hemostasis to proceed via initiation by thromboplastin of the extrinsic pathway. Corn Trypsin Inhibitor is added to the whole blood to inhibit Factor XII and the intrinsic pathway.
Why are there different wall shear rates for the PL and AR chip?
The shear rates are designed to approximate the blood shear rates in-vivo. For the PL chip, platelet aggregation occurs in the arteries and blood shear rates change depending on the size of the artery. In comparison, secondary hemostasis occurs primarily in the venous circulation at much lower shear rates and, again depending on the size of the vein, the in-vivo blood shear rates differ.
Why can't I just use a FX clotting assay?
Presence of a lupus inhibitor or thrombin inhibitor can interfere with the clotting assay. LA patients can produce antibodies that interfere with the phospholipid-dependent clotting reactions that are part of most PT assays. In contrast, the chromogenic factor X reaction does not require a phospholipid membrane surface, and the presence of an LA or DTI does not affect the chromogenic factor X assay.
Why is pNA the leaving group on all of the Chromogenix substrates?
A good chromophore must be readily cleaved by and dissociated from the enzyme. The color must be strong to allow detection of low enzyme activities, but should not interfere with the color of other reactants or impurities in the reaction mixture. It should be water soluble and have low toxicity. The chromophore para-nitroaniline (pNA) fulfills most of these requirements. It is therefore the most common choice of chromophore.
Products: S820316, S820324, S820332, S820340, S820852, S821090, S821413, S822254
Why is thrombin added in the Chromogenix Coamatic® FVIII factor reagent?
Won't tPA be inhibited by PAI-1?
tPA is inhibited in vitro by plasminogen activator inhibitor, so something must be done to avoid this. Acidification of whole blood is therefore performed immediately after withdrawal. This can be done by mixing 1 ml of the citrated blood with 1 ml acetate buffer.