Chromogenix S-2238™
$0.00
Chromogenix S-2238™ is a chromogenic substrate for thrombin. The substrate has been used for the determination of:
- Prothrombin in plasma
- Antithrombin in plasma
- Platelet factor 3 in plasma
- Heparin in plasma
Each vial contains chromogenic substrate S-2238™ 25 mg and mannitol 120 mg as a bulking agent.
Stability
- Substance: Stable until expiry date if stored at 2-8°C. Avoid exposure to light. The substance is hygroscopic and should be stored dry.
- Solution: 1 mmol/L in H2O is stable for more than 6 months at 2-8°C.
| Chemical name | H-D-Phenylalanyl-L-pipecolyl-Larginine-p-nitroaniline dihydrochloride. |
| Formula | H-D-Phe-Pip-Arg-pNA·2 HCl |
| Mol. wt | 625.6 |
| e316 nm | 1.27 . 104 mol-1 . L . cm-1 |
| Solubility | > 10 mmol/L in H2O |
| Suitable stock solution | 1-2 mmol/L in H2O. |
Downloads
- Package Insert (PDF)
- Safety Data Sheet (PDF)
- CofA - Lot(s): N0542548 - Exp: 05/31/2027 (PDF)
- CofA - Exp: 11/29/2027 (PDF)
- CofA - Exp: 04/30/2028 (PDF)
- CofA - Exp: 04/30/2028 (PDF)
- CofA - Exp: 09/30/2028 (PDF)
- CofA - Exp: 09/30/2028 (PDF)
- CofA - Exp: 09/30/2028 (PDF)
- Brochure (PDF)
- Flyer (PDF)
- "Count on Diapharma" Anticoagulation Measurement Flyer (PDF)
- Immunoglobulin Testing Assays Flyer (PDF)
The majority of the Chromogenix substrate library has an Arginine (Arg or R) group at the P1 position (the amino acid position that occurs at the preferred cleavage site). Why is this?
The Chromogenix line is geared toward the proteins involved in hemostasis. These are a group of proteolytic enzymes that comprise the serine proteases, which cleave mainly at the C-terminal side of the basic amino acids arginine or lysine. The peptides at the P2, P3, and P4 positions contribute to the substrate’s specificity. Note that the substrates for plasmin cleave at a lysine group. Other protease groups are aspartic proteases (like pepsin), metallo proteases, and cysteine proteases (which include caspases, with an asp cleavage site).
What if the reconstituted substrate has some precipitate or is cloudy?
The substrate solution is usually prepared with sterile water, but sometimes they may not dissolve properly. Sonication may help, or substrates with low solubility in water can be dissolved in DMSO, then diluted in water. The final DMSO concentration should preferably not exceed 10% in the reaction mixture. It should be noted that stability in DMSO is decreased, as it also is with alkaline buffers.
Why is pNA the leaving group on all of the Chromogenix substrates?
A good chromophore must be readily cleaved by and dissociated from the enzyme. The color must be strong to allow detection of low enzyme activities, but should not interfere with the color of other reactants or impurities in the reaction mixture. It should be water soluble and have low toxicity. The chromophore para-nitroaniline (pNA) fulfills most of these requirements. It is therefore the most common choice of chromophore.
Why are the reactions measured at 405 nm?
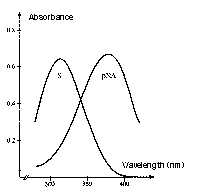
The absorption intensity is expressed by the Beer-Lambert law, A = e x c x l, where A is absorbance, c is molar concentration, l is the path length of sample cell (usually 1 cm), and e is the extinction or molar absorptivity coefficient. The absorption spectrum of the substrate versus the chromophore, pNA (the chromogenic leaving group) is the reason for reading the reactions at 405 nm. The absorbance maximum of the unhydrolyzed, intact substrate is 316 nm and 380 nm for pNA. Although the difference between substrate and product is maximal at 385 nm, at 405 nm, there is less background reading, and the absorbance of the substrate is still less than 1% of that of an equimolar amount of pNA.
What aspects must a scientist consider when choosing the best chromogenic substrate?
Synthetic substrates are very sensitive; they can detect very low enzyme activities and are often more sensitive than a corresponding natural substrate. On the other hand, they can be less selective, or, have less discrimination in their reactivities toward related enzymes compared to the natural substrate. There are steps a scientist can take to maximize sensitivity and specificity. If the specificity of the enzymatic activity to be measured is known then a substrate selectivity table which shows the cross-reactivity of the substrates with different enzymes, and the kinetic data, such as that provided in the Chromogenix catalog, can be helpful. If the specificity of the enzyme is unknown, a screening procedure can be applied. This involves comparing the rate of hydrolysis obtained with different substrates. The presence of contaminating enzymes must also be taken into account. To eliminate interference, an inhibitor can be introduced, the sample can be further diluted, or conditions can be found where the relative activities are optimized. For instance, S-2222™ is selective for FXa, but also for trypsin. If a researcher wants to measure FXa, s/he can add an inhibitor to trypsin, such as soybean trypsin inhibitor. Temperature, pH, buffers, and ionic strength can all affect the rate of hydrolysis and must be considered. Substrate concentration is also important, and a concentration of 2 x Km is usually appropriate. A good substrate has a low Km, meaning maximum reaction velocity is achieved at a low substrate concentration. In other words, the enzyme has a high affinity for the substrate. A high Kcat is also desired, which means the enzyme has a high turnover rate with the substrate (fast reaction).
Define a katal.
One katal (kat) is the amount of enzyme that converts one mole of substrate per second. Activated enzymes from Chromogenix such as FXa and thrombin are measured in nkat. 1 nkat = 1 x 10-9 mol of product released per second.
What is a chromogenic substrate composed of?
Chromogenic substrates are peptides that react with proteolytic enzymes under the formation of color. Chromogenic substrates are made up of a protecting group, amino acid residue(s), side-chain modification if applicable, and the chromophore. The stereochemistry of some substrates may be designated. For example, in the Chromogenix substrate S-2222™, the protecting group is a benzoyl group, the amino acid residue is Ile (isoleucine – a non-polar hydrophobic amino acid), the side chain modification is Glu(g-OR)- where R is 50% H (hydrogen) and CH3 (methyl group). The P2 and P1 amino acid residues are Gly and Arg, respectively, and the chromophore is pNA (para-nitroaniline).
Define an enzyme and a substrate.
Enzymes are proteins that catalyze most of the chemical reactions that take place in the body. The chemical compound upon which the enzyme exerts its catalytic activity is called a substrate. Proteolytic enzymes degrade their substrates, proteins and peptides, by hydrolyzing one or more peptide bond(s). For information on enzyme kinetics, see the Chromogenix Catalog or contact Diapharma Group, Inc. at info@diapharma.com.
What is a peptide? What is the difference between a tripeptide and a tetrapeptide? How are amino acids linked to form peptides?
A peptide is the name assigned to short polymers of amino acids. They are classified by the number of amino acid units in the chain, called amino acid residues. Tripeptides have three amino acid residues while tetrapeptides have four. A polypeptide is formed when the chain of amino acid residues exceeds several dozen in length. A protein is a molecule composed of one or more polypeptide chains.
Proteins are unbranched polymers of amino acids linked head to tail from carboxyl group to amino group, through formation of covalent peptide bonds. The peptide backbone of a protein consists of the repeated sequence.
-N-Ca– C, where N represents the amide nitrogen, Ca represents the a-carbon atom of an amino acid in the polymer chain, and the final C is the carbonyl carbon of the amino acid. This C is in turn linked to the amide N of the next amino acid, and so on down the line. The unbranched polypeptide chain has two ends, an amino-terminal or N-terminal end and a carboxyl-terminal or C-terminal end.
What substrate research methods are available?
- Chromogenix Chromogenic Substrate S-2222 Trypsin
- Chromogenix Chromogenic Substrate S-2222 Tissue Factor Pathway Inhibitor, type – I (TFPI)
- Chromogenix Chromogenic Substrate S-2238 Antithrombin
- Chromogenix Chromogenic Substrate S-2238 Antithrombin (anti-FIIa)
- Chromogenix Chromogenic Substrate S-2288 Proteolytic activity
- Chromogenix Chromogenic Substrate S-2238 Thrombin
- Chromogenix Chromogenic Substrate S-2228 Tissue Plasminogen Activator (t-PA)
- Chromogenix Chromogenic Substrate S-2251 Plasmin
- Chromogenix Chromogenic Substrate S-2302 Kallikrein
- Chromogenix Chromogenic Substrate S-2302 Kallikrein Inhibitor
- Chromogenix Chromogenic Substrate S-2302 Prekallikrein
- Chromogenix Chromogenic Substrate S-2366 Hirudin
- Chromogenix Chromogenic Substrate S-2765 Factor X
- Diapharma Chromogenic Substrate CS GK (substitute for discontinued Chromogenix S-2266) Kallikrein in urine
- Diapharma Chromogenic Substrate CS UK (substitute for discontinued Chromogenix S-2444) Urokinase
- Diapharma Chromogenic Substrate CS PSA (KLK3) (substitute for discontinued Chromogenix S-2586) Chymotrypsin
Historically, the development of a new chromogenic substrate for a specific protease has always been accompanied by the release of method sheets where the application and the methodology for a particular use were described in detail. Some methods are simple chromogenic assays where a buffer and the substrate are the only reagents to be used (i.e. proteolytic activity). Other methods instead require the use of additional compounds, which are or have been commercially available from Chromogenix or elsewhere, and consist of more reaction steps. These protocols were validated in laboratories according to the equipment and the reagents available at the time. In several cases they have been adopted in research, quality control, or routine laboratories and some of them later became Chromogenix kits now present in our product range.
During the last 20 years, the so-called Method Sheets have been taken as the starting point by several scientists, for the development of assays for particular applications, or studies. In some cases the experimental conditions have been changed according to the particular need of the investigation being done. These methods are now presented in a different form: “Research Methods”. The intention here is to provide assay protocols that are not available as kits, but complementary to our product range. In the following list, you can find the methods developed by Chromogenix for several assays. They have to be considered as general guidelines or basic tools for the development of your own assays, some of them requiring a validation within your laboratory with respect to the reagents and equipment used.
For each method, you can find an updated Bibliography with references, where the method has been used, like as originally described or with modifications. This information should facilitate and accelerate the development of the best test protocol. If you do not have that specific journal issue in your laboratory, you can visit MEDLINE. You can search for particular articles, print the abstract and order the original copy through LOANSOME DOC (and receive the document through your local library). At the same time, on our web site you have the possibility to be updated on the new products from Chromogenix.
Theoretical basis for calculation
The hydrolysis of the chromogenic peptide substrate by the proteolytic enzyme follows in general the Michaelis-Menten kinetics. This means that, if the substrate is present at a sufficiently high concentration or if a comparatively small fraction of the substrate is hydrolized, the rate of product (color) formation is proportional to the activity of the enzyme. The rate of pNA formation, i.e. the increase in absorbance per second, is measured photometrically at 405 nm. At this wavelength the extinction coefficient of pNA is
9600 mol -1 • l • cm -1
The enzymatic activity can be quantified in two ways:
- By comparing the activity of an enzyme with that of a standard preparation, which is defined in terms of a specified number of units set by an international or national authority or society (WHO, NIH etc.), or by the activity present in 1 ml of activated pooled normal plasma (Plasma Equivalent Unit = PEU). The standardization is performed by using at standard curve obtained with at least five different concentrations, each performed in duplicate. The standard material must be of the same kind and of the same quality as the sample which is to be measured. This may be still more important for a secondary or domestic standard.
- By measuring the amount (mol) of substrate split, or rather product formed per unit time (absolute activity).
One unit of enzymatic activity, katal (kat) is defined as the amount of activity that converts one mole of substrate per second under standardized conditions. Such conditions as type of substrate, substrate concentration, buffer, pH, ionic strength and temperature should be given along with unit.
Thus, 1 nkat gives a conversion rate of:
1 x 10-9 mol/sec = 60 x 10-9 mol/min
If the total (measuring) volume used is V (ml), the increase in concentration per minute caused by 1 nkat is
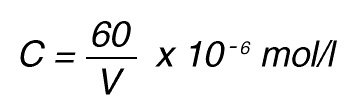
If the absorbance is measured at 405 nm, in a 1 cm cuvette the difference in extinction coefficient is
e = 9600 mol-1 • l
The increase in absorbance/min can then be calculated by using Lambert-Beer’s law:
A = e x C
Thus, 1 nkat gives:
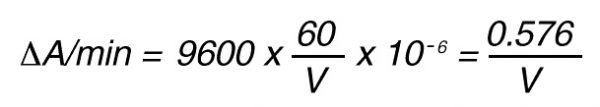
or
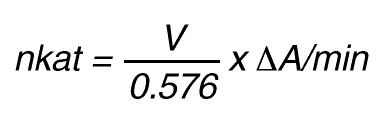
By using a sample volume v (ml):
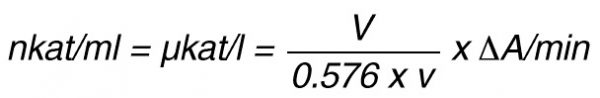
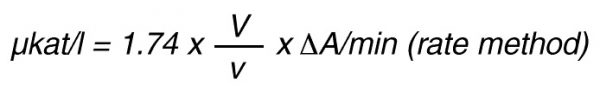
For the end-point method, the incubation time t (min) with substrate is taken into account by the following formula:
According to nomenclature, one unit (U) is the amount of enzyme activity that converts one mol of substrate per minute under standardized conditions. By using the above formulas the units are:
Protein concentrations in plasma
| Component | Molecular Weight kDa | Plasma Concentration mg/l | Plasma Concentration μmol/l |
|---|---|---|---|
| Fibrinogen | 330 | 3000 | 9 |
| Prothrombin | 72 | 150 | 2 |
| Factor V | 330 | 20 | 0.05 |
| Factor VII | 50 | 0.5 | 0.01 |
| Factor VIII | 330 | 0.1 | 0.0003 |
| Factor IX | 56 | 5 | 0.09 |
| Factor X | 59 | 8 | 0.13 |
| Factor XI | 160 | 5 | 0.03 |
| Factor XII | 80 | 30 | 0.4 |
| Factor XIII | 320 | 10 | 0.03 |
| Protein C | 62 | 4 | 0.06 |
| Protein S | 70 | 10 (free) | 0.14 |
| Protein Z | 62 | 2 | 0.03 |
| Prekallikrein | 86 | 50 | 0.6 |
| HMW kininogen | 120 | 70 | 0.6 |
| Fibronectin | 450 | 300 | 0.7 |
| Plasminogen | 92 | 200 | 2 |
| t-PA | 60 | 0.005 | 0.0001 |
| Urokinase | 53 | 0.004 | 0.0001 |
| Antithrombin | 58 | 145 | 2.5 |
| Heparin Cofactor II | 66 | 80 | 1.2 |
| Plasmin Inhibitor | 63 | 60 | 1 |
| Protein C Inhibitor | 57 | 4 | 0.07 |
| α2-Macroglobulin | 725 | 2000 | 3 |
Thrombin IU and Enzyme Activity
The current International Standard for thrombin is the Human a-thrombin 89/588 available from NIBSC. This is a high purity preparation of a-thrombin prepared from Cohn fraction III and assayed by a clotting time method against the first International Standard for thrombin, 75/157.
The National Institute of Health standard (Lot J) is also commonly used for calibration and a study conducted by Gaffney PJ et al. was focused on the relationship between the two standards, and between the International Units and the NIH Units. As a result of this study, based both on a clotting and a chromogenic assay (with the chromogenic substrate S-2238™), 1 NIH-U corresponds to 1.15 IU.
In an article it was shown that bovine thrombin has a higher amidolytic activity than human thrombin when the same NIH-U are compared. It was also underlined that the influence of b and g-forms, that were probably contaminating the bovine enzyme, might be the reason for this discrepancy.
In the same article it was concluded that 1 NIH-U bovine thrombin was equivalent to 3.4 nkat chromogenic substrate S-2238™, and that 1 NIH-U of human thrombin was equivalent to 2.7 nkat chromogenic substrate S-2238™.
From an earlier publication 1 NIH-U of human thrombin corresponded to 2.5 nkat chromogenic substrate S-2238™. The correspondence between NIH-U or IU of thrombin and the enzyme activity expressed in nkat, depends on the substrate, the enzyme preparation (content of a-, b- and g-thrombin) and the assay conditions.
From the article of Friberger , 1 µg thrombin corresponds to 2.2 NIH-U or 5.5 nkat chromogenic substrate S-2238™ or to 0.02 plasma equivalent units. In another study , 1 µg thrombin corresponds to 3.1 NIH-U.
In the experiments done by Chromogenix 1 µg thrombin was equivalent to 3 nkat chromogenic substrate S-2238™ (human) or 4.4 nkat chromogenic substrate S-2238™ (bovine). It might also be added that if all prothrombin is activated in 1 ml of human plasma, about 1.5 nanomoles or 17.5 NIH-U of thrombin are formed.
Thrombin Method
DETERMINATION OF THROMBIN IN PLASMA WITH CHROMOGENIC SUBSTRATE S-2238™
Reagents
- Chromogenic Substrate S-2238™: (0.56 mM) 25 mg Art. No. S820324. Disolve 25 mg lyophlized substrate in 7.14 ml sterile water = 5.6 mM stock solution.
- Buffer: 50 nM Tris, | 0.15, pH 8.3 with 0.2% BSA (10 ml buffer + 100 µl 20% BSA)
Assay
Dilute to a suitable sample concentration (0.5 – 1.0 nkat / ml after dilution with buffer)
| Sample of buffer | 100 µl |
| Incubate at 37°C | 4 min |
| S-2238™ | 100 µl |
| Incubate at 37°C | 3 min |
| Acetic Acid, 20% | 50 µl |
Use buffer as a blank and subtract from sample absorbances
Antithrombin Method
DETERMINATION OF ANTITHROMBIN ACTIVITY IN PLASMA WITH CHROMOGENIC SUBSTRATE S-2238™
Measurement Principle
The antithrombin activity in plasma is measured after addition of an excess of heparin, to form an AT•Heparin complex. An excess of thrombin is then added and allowed to react quantitatively in a 1:1 stoichiometric relationship with the AT Heparin complex present. The residual thrombin splits off p-nitroaniline (pNA) from the chromogenic substrate H-D-Phe-Pip-Arg-pNA (S-2238™). The rate at which pNA is released is measured photometrically at 405 nm. This can be followed on a recorder (initial rate method) or read after stopping the hydrolysis with acid (acid stopped method). The correlation between the change in absorbance per minute (ΔA/min) or absorbance (A) and the AT activity is linear and inversely proportional in the 5-125% range of normal plasma.
| AT + Heparin (excess) | —> | [AT · Heparin] |
| [AT · Heparin] + Thrombin (excess) | —> | [AT · Heparin · Thrombin] +Thrombin (residual) |
| H-D-Phe-Pip-Arg-pNA + H2O | Thrombin (residual) —> |
H-D-Phe-Pip-Arg-OH + pNA |
Reagents
- Chromogenic Substrate S-2238™, 25 mg Art. No. S820324
Reconstitute the substrate S-2238™ (MW: 625.6) in 53 ml of distilled water. Note: Polybrene® can be added to the chromogenic substrate solution at a final concentration of 0.33 mg/ml. - Thrombin, 53 nkat, Art. No. DPGBT-10
Reconstitute with 1.5 ml sterile water. The solution is stable for 4 weeks at 2-8°C. - Buffer – Tris/Heparin, pH 8.4 (25°C)
- Tris 6.1 g (50 mmol/l)
NaCl 10.2 g (175 mmol/l)
Na2EDTA-2H2O 2.8 g (7.5 mmol/l)
Distilled water 800 ml
Adjust the pH to 8.4 at 25°C by adding an appropriate amount (approx. 22 ml) of 1 mol/l HCl. Add 3000 IU of heparin. Fill up to 1000 ml with distilled water. The buffer, if not contaminated, will remain stable for two months at 2-8°C. - Acetic acid 20%
Acetic acid is used in the acid-stopped method.
Specimen collection
Nine parts of freshly drawn venous blood are collected into one part trisodium citrate.
Centrifugation: 2000 x g for 10-20 min at 20-25°C.
Standard curve
Normal plasma has an antithrombin activity of 100%. Two standards (e.g. 25% and 100%) made up fresh should be included in each test run. Check whether ΔA/min or A for the two standards correspond with the stored standard curve. The tolerance limit is ± 0.1 absorbance units. Prepare the standards according to the table below:
| Antithrombin % | Normal plasma ml | Tris/Heparin buffer ml |
| 0 | – | 400 |
| 25 | 100 | 300 |
| 50 | 200 | 200 |
| 75 | 300 | 100 |
| 100 | 400 | – |
Method
Dilute samples and standards as follows:
Tris/Heparin Buffer: 3000 µl
Test plasma or standard: 50 µl
| Initial rate method | |
| Diluted test plasma or standard | 400 µl |
| Incubate at 37°C | 3-6 min |
| Thrombin (20-25°C) | 100 µl |
| Mix and incubate at 37°C | 30 sec |
| Substrate | 300 µl |
Transfer immediately to a 1 cm semi-microcuvette (preheated to 37°C) for measurement of the absorbance change in a photometer at 405 nm and at 37°C, calculate ΔA/min.
| Acid stopped method | |
| Diluted test plasma or standard | 400 µl |
| Incubate at 37°C | 3-6 min |
| Thrombin (20-25°C) | 100 µl |
| Mix and incubate at 37°C | 30 sec |
| Substrate | 300 µl |
| Incubate at 37°C | 30 sec |
| Acetic acid 20% | 300 µl |
Read the absorbance (A) of the sample against distilled water at 405 nm within 4 hours.
Limitations of the procedure
In some pathological states (DIC, sepsis) plasma alone may hydrolyse the chromogenic substrate S-2238™. This interfering reaction may be determined by assay of a test sample in the absence of added thrombin. This activity rarely corresponds to more than 1% of that of the added thrombin.To improve the validity of the assay the value obtained in the absence of added thrombin can be subtracted from the sample value. Bilirubin, haemoglobin and plasma from hyperlipaemic patients interfere in absorbance reading. Patients plasma blanks are necessary in these instances for the acid stopped method only. At concentrations below 25% AT it is recommended to double the plasma concentration (100 µl plasma + 3 ml buffer). The result is then divided by two.
Calculation
Plot A or ΔA/min for the standards against their known antithrombin activity.
Percent of normal AT activity is determined by plotting the A or ΔA/min for the test sample on the standard curve and reading the corresponding AT value.
Bibliography
- Odegard OR et al. Heparin cofactor activity measured with an amydolytic method. Thromb Res 6, 287-294 (1975).
- Odegard OR et al. Evaluation of an amidolytic heparin cofactor method. Thromb Res 7, 351-360 (1975).
- Abildgaard U et al. Antithrombin (heparin cofactor) assay with new chromogenic substrates. Thromb Res 11, 549-553 (1977).
- Kahlé LH et al. Antithrombin III, Evaluation of an automated antithrombin III method. Thromb Res 12, 1003-1014 (1975).
Heparin Method
DETERMINATION OF HEPARIN IN PLASMA WITH CHROMOGENIC SUBSTRATE S-2238™
Measurement Principle
Heparin is analysed as a complex with antithrombin (AT) present in the sample. The concentration of this complex is dependent on the availability of AT. In order to obtain a more constant concentration of AT, purified AT is added to the test plasma. Thrombin in excess is neutralized in proportion to the amount of heparin, which determines the amount of heparin-AT complex. The remaining amount of thrombin hydrolyses the chromogenic substrate H-D-Phe-Pip-Arg-pNA (Chromogenic Substrate S-2238™) thus liberating the chromophoric group, pNA. The color is then read photometrically at 405 nm.
| Heparin + AT | —> | [Heparin · AT] |
| [Heparin · AT] + Thrombin (excess) | —> | [Heparin · AT · Thrombin] + Thrombin (residual) |
| H-D-Phe-Pip-Arg-pNA + H2O | Thrombin (residual) —> |
H-D-Phe-Pip-Arg-OH + pNA |
Reagents
- Chromogenic Substrate S-2238™, 25 mg Art. No. S820324
Reconstitute the substrate S-2238™ (MW: 625.6) with 40 ml of distilled water. - Thrombin
Human thrombin or bovine thrombin can be used in 0.15 mol/l NaCl solution.
The activity of the solution should be 14 nkat/ml (about 6 NIH-U/ml).
If bovine thrombin 53 nkat from Diapharma (Art. No. DPGBT-10) is used, dissolve the content of one vial with 3.8 ml saline. - Antithrombin 10 IU Art No. B820720
Reconstitute with 5 ml water to obtain a concentration of 2 IU/ml. - Tris Buffer, pH 8.4 (25°C)
Tris 6.1 g (50 mmol/l)
NaCl 10.2 g (175 mmol/l)
Na2EDTA 2.8 g (7.5 mmol/l)
Distilled water 800 ml
Adjust the pH to 8.4 at 25°C by adding an appropriate amount (approx. 22 ml) of 1 mol/l HCl. Fill up to 1 liter. - Normal plasma
Blood should be taken from normal donors. 10-30 ml of citrated blood (9 vol blood and 1 vol 0.1 mol/l sodium citrate) are taken from each donor. The first ml of blood is discarded and the tube is kept in an ice bath. Plasma is prepared by centrifugation at 2000 x g for 20 minutes at 4°C. Equal amounts of plasma from the donors are mixed and dispensed in small volumes. The normal plasma is stable for 3 months at -20°C or below. Thaw at 37°C and then keep on ice. - Acetic acid 20%
Acetic acid is used in the acid-stopped method.
Specimen collection
Blood (9 vol) is mixed with sodium citrate (1 vol) cooled to 0°C with ice and centrifuged at 2000 x g for 20 min at 4°C.
Dilute plasma 1:5 with Tris Buffer pH 8.4.
Standard curve
The same heparin as is used for the patient is diluted to 1 IU/ml with saline 0.9%. Then 100 µl dilution is further diluted with 1.9 ml buffer to obtain a concentration of 0.05 IU/ml.
| Standard IU/ml |
Buffer ml |
AT ml |
Plasma dil 1:5 ml |
Heparin 0.05 IU/ml ml |
| 0.00 | 800 | 100 | 100 | 0 |
| 0.25 | 700 | 100 | 100 | 100 |
| 0.50 | 600 | 100 | 100 | 200 |
| 0.75 | 500 | 100 | 100 | 300 |
| 0.10 | 400 | 100 | 100 | 400 |
Method
| Initial rate method | Tube No. 1 |
| Buffer | 800 µl |
| AT | 100 µl |
| Test plasma | 100 µl |
| Mix | |
| Tube No. 2 | |
| Standard or tube No. 1 | 200 µl |
| Incubate at 37°C | 3-4 min |
| Thrombin | 100 µl |
| Incubate at 37°C | 30 sec |
| Substrate (37°C) | 200 µl |
| Mix |
Transfer sample immediately to a 1 cm micro-cuvette (preheated at 37°C) for measurement of the absorbance change at 405 nm. Calculate ΔA/min. Read the absorbance against a normal plasma blank in a photometer at 405 nm.
| Acid stopped method | Tube No. 1 |
| Buffer | 100 µl |
| AT | 100 µl |
| Test plasma | 100 µl |
| Mix | |
| Tube No. 2 | |
| Standard or tube No. 1 | 200 µl |
| Incubate at 37°C | 3-4 min |
| Thrombin | 100 µl |
| Incubate at 37°C | 30 sec |
| Substrate (37°C) | 200 µl |
| Incubate at 37°C | 60 sec |
| Acetic acid 20% | 300 µl |
| Blanks for acid stopped method | Normal plasma blank | Test plasma blank |
| Standard 0 IU/ml | 200 µl | – |
| Sample from tube No. 1 | – | 200 µl |
| Acetic acid | 300 µl | 300 µl |
| Mix | ||
| Distilled water | 300 µl | 300 µl |
| Mix |
Note: As a rule a normal plasma blank or even water is used as a blank. If bilirubin exceeds 100 mmol/l or the test plasma is opaque, read the test plasma sample against its own blank.
Calculation
Plot A or ΔA/min for the standards against their known heparin concentration.
Heparin concentration is determined by plotting the A or ΔA/min for the test sample on the standard curve and read the corresponding heparin value.
Bibliography
- Bhargava AS et al. Characterization of a new potent heparin. 2nd communication: chemical analysis of the carbohydrate content and determination of the biological activity of a new potent heparin preparation in vitro, using protamine nutralization and amidolytic methods for factor Xa and thrombin. Arzneimittelforschung 30, 1071-1074 (1980).
- Sache E et al. Studies on a highly active anticoagulant fraction of high molecular weight isolated from porcine sodium heparin. Thromb Res 25, 443-458 (1982).
- Van Putten J et al. Determination of low molecular weight heparin in clinical laboratory. Haemostasis 14, 205-210 (1984).
- Van Putten J et al. Automated spectrophotometric heparin assays. Comparison of methods. Haemostasis 14, 195-204 (1984).
- Berry CN et al. Effects of the synthetic thrombin inhibitor argatroban on fibrin- or clot-incorporated thrombin: comparison with heparin and recombinant hirudin. Thromb Haemost 72, 381-386 (1994).
- Byun Y et al. Effect of fibronectin on the binding of antithrombin III to immobilized heparin. J Biomed Mater Res 30, 95-100 (1996).
Prothrombin Activity Method
DETERMINATION OF PROTHOMBIN ACTIVITY IN PLASMA WITH CHROMOGENIX S-2238™
Background
A number of studies during the last few years support the notion that venous thromboembolism (VTE) is a multifactorial disease most often triggered by circumstantial risk factors (trauma, surgery, pregnancy, oral contraceptives, immobilization and age) in combination with one or more genetic or acquired coagulation disorders (see ref. 1 of a review).
Elevated activity of prothrombin in the absence of a known underlying genetic disorder is also associated with an increased thrombotic risk2.
A mutation G → A in the untranslated 3’-region of the prothrombin gene at nucleotide position 20210 constitutes a risk factor for VTE with an odds ratio of 3-52-10. About 90% of the carriers of this mutation have elevated levels (>115%) of prothrombin activity2,7,8. Levels above the upper limit of the normal range (75 – 130%) are commonly hetero- and homozygotes2,7-9.
So far, there is no explanation why a comparatively mild increase of prothrombin activity constitutes a risk factor for thrombosis and this is therefore an area of active clinical and biochemical research. Chromogenic methods for accurate determination of elevated activities of prothrombin and other coagulation factors, such as factor VIII11,12 are important tools for assessing the risk for VTE in patients and family members.
Measurement Principle
Prothrombin is activated to meizothrombin by the snake venom enzyme Ecarin from Echis Carinatus.
After a certain incubation time, the amount of meizothrombin formed is measured with the thrombin selective substrate Chromogenix S-2238™, which also is cleaved by meizothrombin.
The absorbance recorded at 405nm is proportional to the prothrombin activity in the sample.
| Prothrombin | Ecarin —> |
Meizothrombin |
| S-2238™ | Meizothrombin —> |
pNA + Peptide |
Reagents
- Tris BSA Buffer (catalog# TB031-20)Buffer for sample dilution, containing 0.5mol/l Tris HCl pH 7.3, I = 2.0 with NaCl and 2% bovine serum albumin. Before use, dilute the stock solution 1+9 with sterile water to obtain a buffer working solution. The buffer working solution is prepared and used within the same day.
- Ecarin Diluent (catalog# ED0413-20)Buffer for dilution of Ecarin, containing 0.05mol/l Tris HCl pH 7.6, I=0.15 with NaCl, bovine serum albumin, polyethylene glycol and a fibrin polymerization inhibitor.
- Ecarin (catalog# ECARIN50B)Reconstitute with sterile water according to the Ecarin package insert. Freeze in suitable aliquots at -20°C or at -70°C. Stable for 3 months at both storage temperatures. Before use, dilute with Prothrombin Activator Diluent to obtain a concentration of 2.4U/ml. Stable for 8 hours at 20-25°C and for 1 week at 2-8°C.Note: Echis Carinatus crude venom can also be used. A suitable final concentration of this reagent is approximately 5µg/ml; however, this may vary between different sources. 10-20% loss of activity may occur upon freezing at -20°C.
- S-2238™ (catalog# S820324)Reconstitute with 13ml of sterile water to obtain a 3mmol/l solution.
Specimen Collection
Blood (9 volumes) is mixed with 0.1mol/l sodium citrate (1 volume) and centrifuged at 2000 x g for 20 min at 20-25°C. Separate plasma carefully from blood cells. Perform the analysis within 24 hours when plasma is stored at 2-25°C. Alternatively, freeze aliquots ≤ 1ml at -20°C or below. Perform the analysis of frozen samples within two months when stored at -20°C or within one year when stored at -70°C or below. No significant loss of prothrombin activity occurs upon freezing once, provided freezing is made in small aliquots (< 1 ml) and thawing is performed in a water bath or in an electric heater at 25-37°C.
Sample and Standard Dilutions
Standards
Calibrated normal plasma is diluted 1:23 – 1:160 to provide standard concentrations of 25-175%. The following table provides a suggestion of standard dilutions.
| Standard Dilution | Prothrombin Activity |
| 1:18 | 167% |
| 1:22 | 136% |
| 1:30 | 100% |
| 1:60 | 50% |
| 1:120 | 25% |
Samples
Plasma samples are diluted 1:40 in Tris BSA Buffer working solution for application on microplate and diluted 1:80 for application on ACL (see below).
Microplate Assay Procedure
| Standard/Sample dilution | 50μl |
| Incubate at 37°C | 2-4min |
| Ecarin or Echis Carinatus (37°C) | 50μl |
| Incubate at 37°C | 3min |
| Substrate (37°C) | 50μl |
| Read kinetically or incubate at 37°C | 2min |
| Acetic acid, 20% | 50μl |
Determine the absorbance difference A405nm-490nm for the standard dilution and the samples. Draw a standard curve from the absorbances obtained for the standard dilutions. Read the prothrombin activity for the samples from the standard curve.
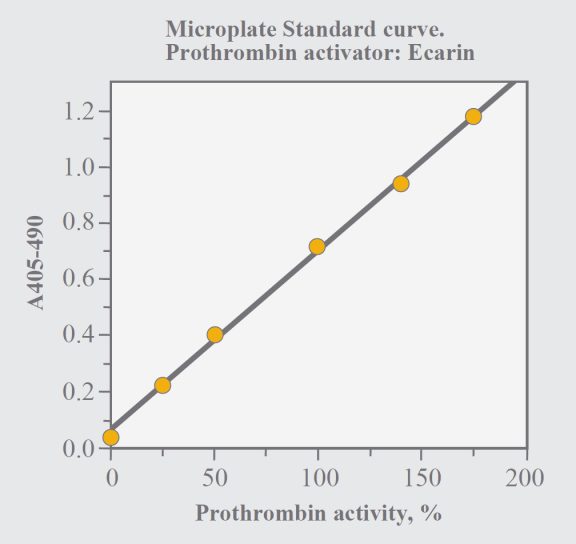
Fig. 1. Standard curve with the microplate method.
Application on ACL
Use the plasminogen channel program. Prepare a standard dilution 1:40, which corresponds to a nominal prothrombin activity of 100% (see above regarding calibration). Standard dilutions corresponding to 25% and 50% are then automatically prepared by the instrument. In order to allow determination of prothrombin activity up to 200%, sample plasma should be diluted 1:80 and the obtained result should be multiplied with two.
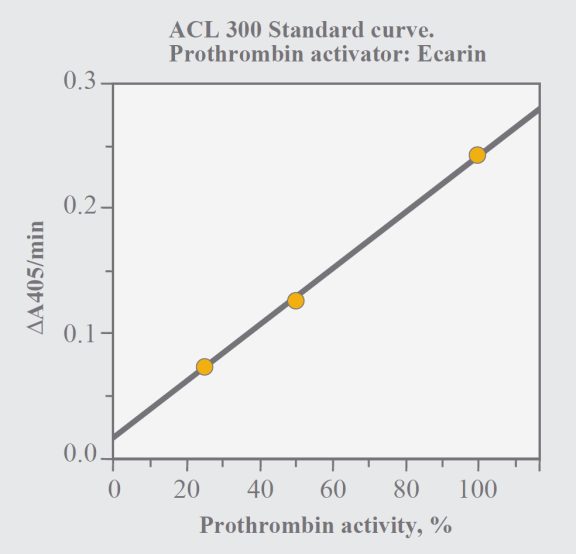
Fig. 2. Standard curve with the ACL method
Expected values13
The normal range is 75 – 130% (mean 102% 2 SD) as determined from analysis in microplate and on the ACL 300 of 101 healthy individuals (49 men and 52 women; age range 20 – 68 years). Analysis of plasma from 42 carriers of the G20210A mutation, who were not on oral anticoagulant treatment at the time of blood sampling, resulted in an activity range of 94 – 164% (mean 128% 2SD).
Interference and Limitations
No influence in the assay is obtained from variation of antithrombin activity in the range 50 – 150% of normal. Since meizothrombin is formed and measured, no influence in the assay is obtained from heparin levels ≤ 1 IU/ml plasma. Since Ecarin also activates decarboxyprothrombin, which is produced during oral anticoagulant therapy with anti-vitamin K drugs, plasma from patients undergoing such treatment should not be analysed with this method.
Repeatability
The imprecision, expressed as CV, within and between series (7 series, 5 replicates in each series) is ≤4% at 50% and 100% prothrombin activity.
Bibliography
- Lane DA, Mannucci PM, Bauer KA, Bertina RM, Bochkov NP, Boulyjenkov V, Chandy M, Dahlbäck B, Ginter EK, Miletich Jp, Roosendaal FR, Selingsohn U.
Inherited Thrombophilia: part 1.
Thromb Haemost 76, 651-662 (1996) - Poort SR, Roosendaal FR, Reitsma PH, Bertina RM. A common genetic variation in the 3’-untranslated region of the prothrombin gene is associated with elevated plasma prothrombin levels and an increase in venous thrombosis.
Blood 88, 3698-3703 (1996) - Hillarp A, Zöller B, Svensson PJ, Dahlbäck B.
The 20210 allele of the prothrombin gene is a common risk factor among Swedish out-patients with verified deep venous thrombosis.
Thromb Haemost 78, 990-992 (1987) - Cumming AM, Keeney S, Salden A, Bhavnani M, Shwe RH, Hay CRM. The prothrombin gene G20210A variant: prevalence in a UK anticoagulant clinic population.
Br J Haematol 98, 353-355 (1997) - Brown K, Luddington R, Williamson D, Baker P, Baglin T.
Risk of venous thromboembolism associated with a G to A transition at position 20210 in the 3’-untranslated region of the prothrombin gene.
Br J Haematol 98, 907-909 (1997) - Makris M, Preston FE, Beuchamp NJ, Hampton KK, Daly ME, Cooper P, Bayliss P, Peake IR. Co-inheritance of the 20210A allele of the prothrombin gene increases the thrombotic risk in subjects with familial thrombophilia.
Thromb Haemost 78, Suppl., 165 (1997) - Ferraresi P, Marchetti G, Legnani C, Cavallari E, Castoldi E, Mascoli F, Ardissino D, Palareti G, Bernardi F. The heterozygous 20210G/A prothrombin genotype is associated with early venous thrombosis in inherited thrombophilias and is not increased in frequency in artery disease.
Arterioscl Thromb Vasc Biol 17, 2418-2422 (1997) - Kapur RK, Mills LA, Spitzer SG, Hultin MB. A prothrombin gene mutation is significantly associated with venous thrombosis.
Arterioscl Thromb Vasc Biol 17, 2875-2879 (1997) - Howard TE, Marusa M, Channel C, Duncan A. A patient homozygous for a mutation in the prothrombin gene 3’-untranslated region associated with massive thrombosis.
Blood Coag Fibrinol 8, 316-319 (1997) - Martinelli I, Sacchi E, Landi G, Taioli E, Duca F, Mannucci PM. High risk of cerebral-vein thrombosis in carriers of a prothrombin-gene mutation and in users of oral contraceptives.
New Engl J Med 338, 1793-1797 (1998) - Koster T, Blann AD, Briët E, Vanenbroucke JP, Roosendaal FR. Role of clotting factor VIII in effect of von Willebrand factor on occurrence of deep-vein thrombosis.
The Lancet 345, 151-155 (1995) - D’Onnel J, Tuddenham EGD, Manning R, Kemball-Cook G, Johnson D, Laffan M. High prevalence of elevated factor VIII levels in patients referred for thrombophilia screening: role of increased synthesis and relationship to the acute phase reaction.
Thromb Haemost 77, 825-828 (1997) - Rosén S, Andersson M, Ghosh R. A new chromogenic prothombin method providing accurate determination of elevated prothombin activity in plasma samples.
ISTH 1999, Abstract 269